Procedure At A Glance
Total Time of IHC Staining: ~4–6 Hours (excluding overnight incubation)
- Deparaffinize: Xylene (3 changes, 10 mins each) → Rehydrate in graded Alcohols (100% to 50%) → Water.
- Antigen Retrieval: Heat slides in Citrate Buffer (pH 6.0) or EDTA (pH 9.0) via pressure cooker/steamer (~20 mins). Cool to room temp.
- Wash: PBS or TBS (3 changes, 5 mins each).
- Block: Apply Blocking Buffer (15–30 mins) to prevent non-specific binding.
- Primary Antibody: Incubate with specific primary antibody (Overnight at 4°C OR 1 hour at Room Temp).
- Wash: PBS/TBS (3 changes).
- Secondary Antibody: Incubate with HRP-conjugated secondary antibody (30–60 mins at Room Temp).
- Wash: PBS/TBS (3 changes).
- Develop: Apply DAB Chromogen (5–10 mins). Watch for brown color change.
- Counterstain: Hematoxylin (1–2 mins) → Rinse → Bluing Reagent.
- Mount: Dehydrate (Alcohols) → Clear (Xylene) → Apply Coverslip.
Introduction
Immunohistochemistry (IHC) staining is a powerful technique that allows us to peek into the intricate world of protein expression within tissues. Immunohistochemistry (IHC) staining transcends the limitations of traditional histological stains like hematoxylin and eosin (H&E) that primarily assess tissue morphology. Immunohistochemistry (IHC) staining allows us to visualize specific proteins in situ, providing invaluable insights into the diagnosis, prognosis, and even treatment of various diseases.
Role and Purpose of Immunohistochemistry (IHC) Staining
- Diagnosing diseases: In hematopathology, immunohistochemistry (IHC) plays a crucial role in differentiating various types of lymphomas, leukemias, and other blood malignancies. Identifying specific protein markers using immunohistochemistry (IHC), such as CD3 for T-cell lymphomas or CD20 for B-cell lymphomas, allows for accurate diagnosis and classification, guiding appropriate treatment strategies.
- Subclassify malignancies: Within the spectrum of lymphomas, immunohistochemistry (IHC) plays a vital role in classifying subtypes. By detecting specific markers like CD20, CD3, and BCL2 using immunohistochemistry (IHC), we can accurately subdivide lymphomas, each with distinct clinical features and prognoses, paving the way for personalized therapy.
- Prognosticating disease course: Immunohistochemistry (IHC) staining can reveal the expression of proteins associated with disease progression or treatment resistance. For instance, detecting elevated Ki-67 expression (a proliferation marker) in a lymphoma sample can indicate a more aggressive disease course.
- Understanding disease mechanisms: By visualizing the spatial distribution of proteins in tissues, immunohistochemistry (IHC) staining sheds light on cellular interactions and biological processes involved in disease development. In the context of hematological disorders, immunohistochemistry (IHC) can help elucidate the role of specific protein pathways in lymphomagenesis or leukemogenesis.
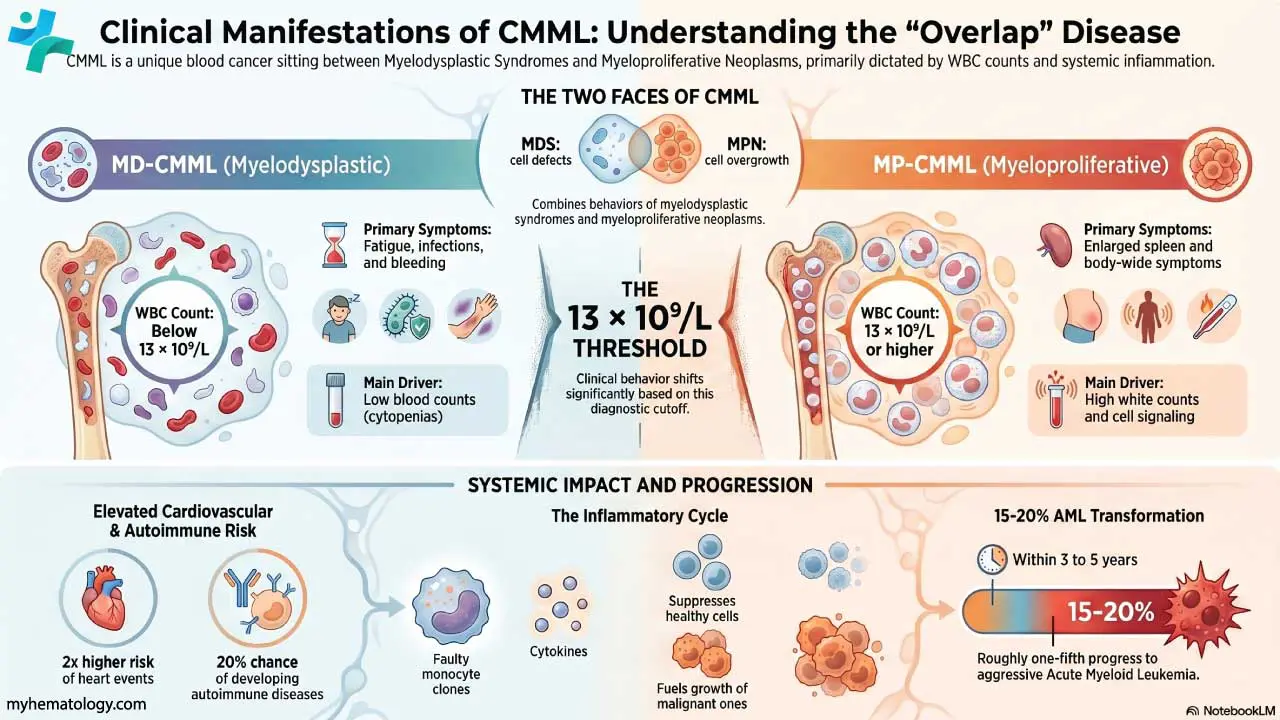
Hematological Disorders
Hematological malignancies, particularly lymphomas, present a prime example of where immunohistochemistry (IHC) shines brightest. These cancers, characterized by abnormal proliferation of lymphocytes, often require specific protein markers for definitive diagnosis and subclassification. Immunohistochemistry (IHC) plays a critical role in identifying these markers, including:
- B-cell lymphomas: Detection of CD20, BCL2, and CD3 is crucial for differentiating B-cell lymphomas from T-cell lymphomas and other types, guiding appropriate treatment strategies.
- T-cell lymphomas: Immunohistochemistry (IHC) markers like CD3, CD4, and CD8 help distinguish various subtypes of T-cell lymphomas, each with distinct clinical courses and treatment needs.
- Hodgkin lymphoma: Identifying specific markers like CD30 and CD15 helps differentiate Hodgkin lymphoma from other lymphoma subtypes, leading to accurate diagnosis and prognosis.
Automated Vs Manual Staining
While manual staining allows for the ultimate control over a single slide, automated systems have become the gold standard in clinical hematology labs to ensure every CD20 or Ki-67 stain looks identical, day after day. Standardization is the biggest win for hematology, as many diagnostic markers (like BCL-2 or Cyclin D1) rely on specific intensity thresholds that are much easier to maintain on a machine.
Manual vs. Automated Comparison
| Feature | Manual Staining | Automated Staining |
| Consistency | Variable; depends on technician skill. | High; strictly standardized protocols. |
| Throughput | Low; labor-intensive for the tech. | High; can process 30–100+ slides at once. |
| Labor Cost | High (Requires constant "hands-on" time). | Low (Technician "loads and walks away"). |
| Initial Setup Cost | Very Low (Jars, reagents, and a timer). | Very High (Expensive machinery). |
| Flexibility | High; easy to tweak steps on the fly. | Low; limited to validated "on-board" protocols. |
| Risk of Error | High (Wrong incubation, slides drying out). | Low (Sensors detect reagent levels). |
Principle of Immunohistochemistry (IHC)
Preparation
Thin slices of tissue are fixed to preserve their structure and then treated to permeabilize cell membranes, allowing antibodies access to their targets. This process, called antigen retrieval, ensures the protein is accessible for its pursuer.
The Antibody Hunt
The primary antibody, Y-shaped molecules designed to recognize and bind to specific proteins or other antigens, tailored to the protein of interest, is applied to the tissue.
Amplifying the Signal
To visualize the binding, we need a secondary antibody linked to a reporter molecule. A secondary antibody, tagged with a fluorescent dye or enzyme, is introduced. This secondary antibody binds to the first antibody-antigen complex.
Visualization using a Reporter Molecule
If a fluorescent dye is used, the dye attached to the secondary antibody emits light of a specific wavelength when excited by a light source. the bound protein glows under a microscope, revealing its location in the tissue
If an enzyme is attached, a substrate is added, triggering a color-changing reaction. The protein's presence becomes evident as a brown, red, or blue stain, marking its hiding place. This can be an enzyme like horseradish peroxidase (HRP). Upon receiving a specific signal, the enzyme converts a colorless substrate into a vibrant brown precipitate, marking the location of the antigen-antibody complex.
Results
Finally, the tissue is mounted and examined under a microscope. By interpreting the brown stains or fluorescent signals, we can pinpoint the distribution and abundance of specific molecules within the tissue, providing valuable insights into disease processes and aiding in diagnosis and treatment.
Antibody Types
In Immunohistochemistry (IHC), the antibodies are the "workhorses" of the diagnostic process. Choosing the right type is the difference between a clear, diagnostic signal and a messy slide.
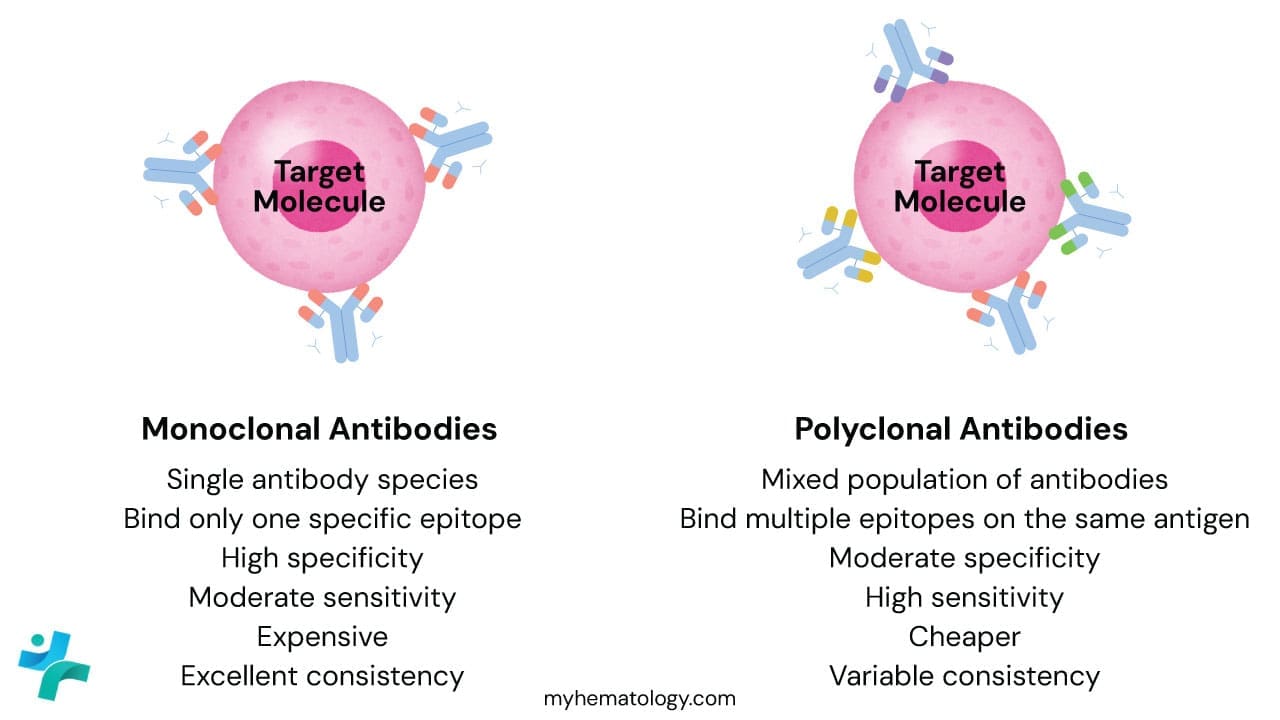
Monoclonal Antibodies
Monoclonal antibodies are produced by a single clone of B-cells (usually via hybridoma technology). Every single antibody molecule in the vial is identical, recognizing the exact same epitope (binding site) on the target protein.
- The Advantage: High Specificity. Because they only "see" one specific spot on a protein, they rarely bind to the wrong things. This leads to very low background staining (cleaner slides).
- The Disadvantage: If that one specific epitope is damaged or masked by formalin fixation, the antibody won't bind at all, leading to a false negative.
- Example: The L26 clone is a famous monoclonal antibody used to detect CD20 in B-cell lymphomas.
Polyclonal Antibodies
Polyclonal antibodies are harvested from the serum of an animal (like a rabbit or goat) that has been immunized with the target protein. It is a mixture of antibodies produced by many different B-cell clones.
- The Advantage: High Sensitivity. These antibodies recognize multiple different epitopes on the same protein. If one binding site is damaged during tissue processing, the others will still catch the target. They are great for detecting proteins expressed at low levels.
- The Disadvantage: High Background. Because there are so many different antibodies in the mix, there is a higher chance one of them will "cross-react" with a similar-looking protein that isn't your target.
- Example: Polyclonal Rabbit anti-Human Myeloperoxidase (MPO) is often used to identify myeloid cells in bone marrow biopsies.
Monoclonal vs. Polyclonal Comparison
| Feature | Monoclonal | Polyclonal |
| Source | Single B-cell clone (Hybridoma) | Multiple B-cell clones (Serum) |
| Epitope Binding | Only one specific epitope | Multiple epitopes on the same antigen |
| Specificity | Very High | Low to Moderate |
| Sensitivity | Moderate | Very High |
| Batch-to-batch Consistency | Excellent (identical every time) | Variable (each batch is slightly different) |
| Cost | More expensive | Generally cheaper |
Application Types: Direct vs. Indirect
Beyond how they are made, we categorize them by how they are applied to the slide:
- Direct IHC: The primary antibody itself is chemically "tagged" with a label (like a fluorescent dye or an enzyme).
- Pros: Fast.
- Cons: Not very sensitive because only one label is attached per antigen.
- Indirect IHC: This is the industry standard. You apply an unlabeled primary antibody first, followed by a "Secondary Antibody" that is tagged with the enzyme.
- Pros: Signal Amplification. Multiple secondary antibodies can bind to a single primary antibody, making the final "brown stain" much darker and easier to see.
Species of Origin
It is also important to note which animal the antibody was raised in (usually Mouse or Rabbit). If your primary antibody was raised in a mouse, your secondary antibody must be an "Anti-Mouse" antibody. If you mix them up, the stain simply won't work.
Materials
This protocol is a general guideline and may need to be adapted based on specific antibodies, tissues, and desired staining methods. Always consult the datasheets and recommendations of the antibody and reagent manufacturers before proceeding.
- Tissue sections on positively charged slides
- Xylene
- Ethanol solutions (100%, 95%, 70%, 50%)
- Deionized water
- Antigen retrieval buffer (specific to your target antigen)
- Blocking buffer (e.g., 1% BSA in PBS or 5% serum from the secondary antibody's host)
- Primary antibody diluted in antibody diluent (concentration varies, refer to datasheet)
- Secondary antibody conjugated to reporter molecule (HRP, fluorescent dye)
- DAB chromogen solution (for HRP detection) or fluorescent mounting medium
- Hematoxylin and eosin (H&E) staining reagents (optional)
- Coplin jars or staining dishes
- Microscope slides and coverslips
- Forceps and lint-free wipes
- Pipettes and pipette tips
- Wash buffer (PBS or TBS)
- Waste disposal containers
Protocol
- Deparaffinization and Rehydration:
- Place slides in a Coplin jar containing xylene for 10 minutes at room temperature (repeat 2-3 times).
- Transfer slides to 100% ethanol for 5 minutes (repeat 2 times).
- Descend through graded ethanol solutions (95%, 70%, 50%) for 2 minutes each.
- Rinse slides briefly in deionized water.
- Antigen Retrieval (Optional): This step may be necessary to unmask hidden epitopes, depending on the antibody and tissue fixation. Refer to the antibody datasheet for specific recommendations.
- Heat the antigen retrieval buffer in a pressure cooker or steamer to the desired temperature and time (follow manufacturer's instructions).
- Place slides in the hot buffer and incubate for the specified time.
- Allow the solution to cool down gradually before removing the slides.
- Rinse slides with deionized water.
- Blocking: Incubate slides in blocking buffer for 15-30 minutes at room temperature to block nonspecific binding sites.
- Primary Antibody Incubation:
- Apply the diluted primary antibody solution to the tissue sections. Cover with a plastic coverslip to prevent drying.
- Incubate in a humidified chamber at the recommended temperature and time (usually overnight at 4°C).
- Washing: Rinse slides gently with wash buffer 3 times for 5 minutes each.
- Secondary Antibody Incubation: Apply the secondary antibody solution diluted in antibody diluent to the tissue sections. Incubate for 30-60 minutes at room temperature in the dark.
- Washing: Repeat step 5.
- Detection:
- HRP detection: Incubate slides in DAB chromogen solution for 5-10 minutes, monitoring for brown precipitate development. Rinse slides with deionized water to stop the reaction.
- Fluorescent detection: Rinse slides with wash buffer and deionized water. Mount slides with fluorescent mounting medium and coverslips.
- Counterstaining and Mounting (Optional):
- Stain slides with hematoxylin and eosin (H&E) for morphological evaluation if desired.
- Dehydrate and clear slides in graded ethanol solutions and xylene.
- Mount slides with permanent mounting medium and coverslips.
- Microscopic Examination: Examine stained slides under a microscope to visualize the specific protein expression patterns revealed by the immunohistochemistry (IHC) staining.
This protocol does not employ a hydrogen peroxide (H₂O₂) block step for several reasons:
- ADEC secondary antibodies: ADEC (antibody-dependent enzyme conjugation) secondary antibodies pre-conjugated to HRP (horseradish peroxidase) is used. These lack the Fc region that interacts with endogenous peroxidase, eliminating the need for H₂O₂ blocking to prevent non-specific staining.
- DAB specificity: The chosen chromogen, DAB (3,3'-diaminobenzidine), reacts specifically with HRP in the presence of H₂O₂. Hence, even if some endogenous peroxidase persists, it wouldn't generate a brown precipitate with DAB, mitigating the need for a block.
- Optimized washing: Thorough washing steps following each antibody incubation effectively remove unbound antibodies and potential endogenous enzymes, further reducing the requirement for an H₂O₂ block.
The decision to exclude H₂O₂ block was based on a combination of factors like antibody type, chromogen choice, and optimized washing steps for the specific tissue and fixation method used.
Interpretation
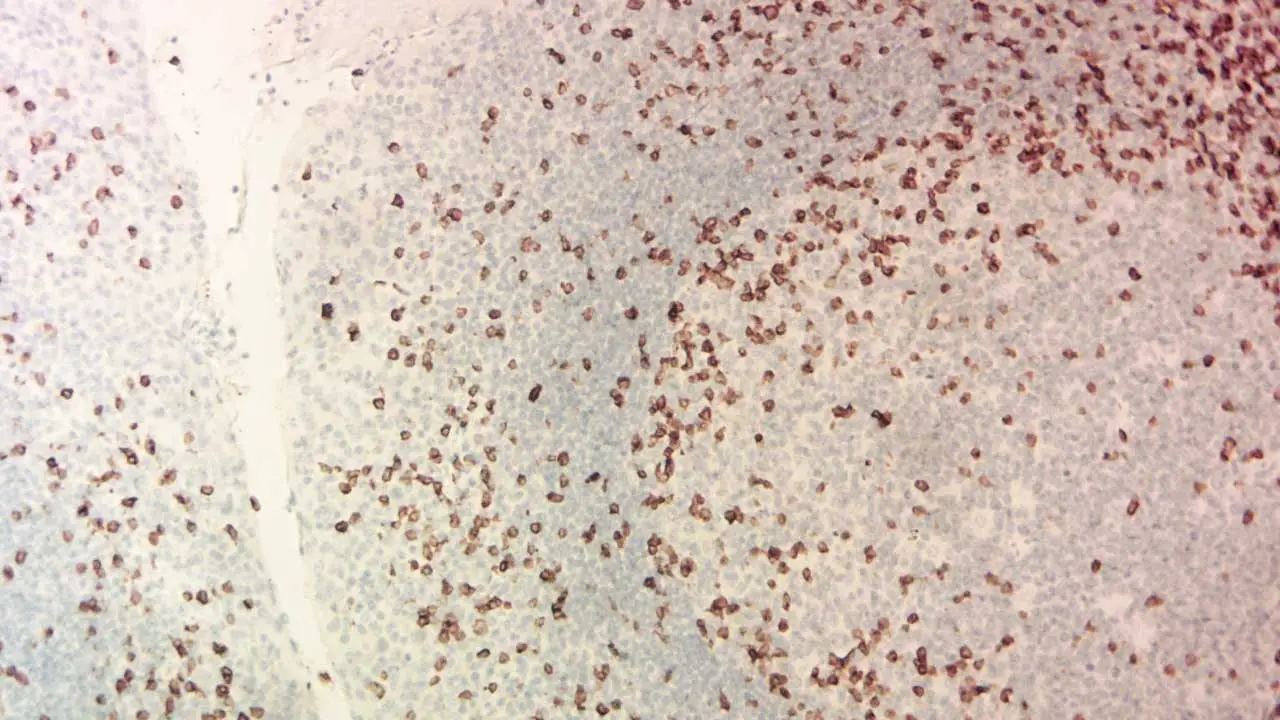
Interpreting immunohistochemistry (IHC) stains requires a keen eye and a thorough understanding of the specific antibody, target antigen, and tissue context.
Staining Patterns
The pattern of antibody binding in immunohistochemistry (IHC) staining reveals valuable information about protein expression and cellular distribution. Some common patterns include:
- Membranous: Staining concentrated on the cell membrane. (Think of a fence marking the cell boundaries)
- Cytoplasmic: Staining filling the cytoplasm within the cell. (Imagine confetti scattered inside the cell)
- Nuclear: Staining focused within the cell nucleus. (Think of a highlighter marking important text in the DNA library)
- Focal: Staining scattered in specific areas or cell clusters. (Like spotlights highlighting certain regions)
- Negative: No detectable staining, indicating the target antigen is absent or below the detection threshold.
Staining Intensity
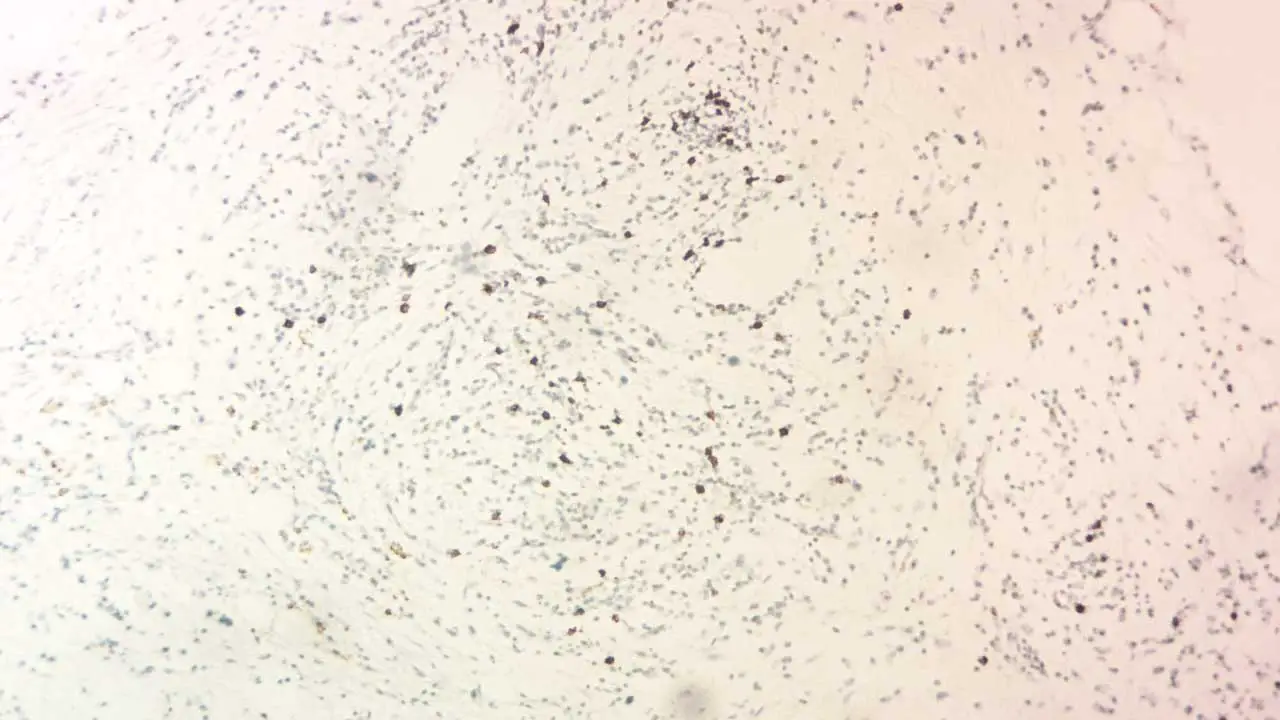
The intensity of the immunohistochemistry (IHC) stain reflects the relative abundance of the target antigen. Different shades can be categorized as:
- Negative: No visible staining.
- Weak: Faint brown or fluorescent signal.
- Moderate: Clear brown or brighter fluorescent signal.
- Strong: Intense brown or highly fluorescent signal.
By combining the staining pattern and intensity, we can formulate a preliminary interpretation of the immunohistochemistry (IHC) results. However, remember that definitive diagnosis and conclusions often require correlation with other clinical findings and pathological examinations.
Positive vs. Negative Controls in IHC staining
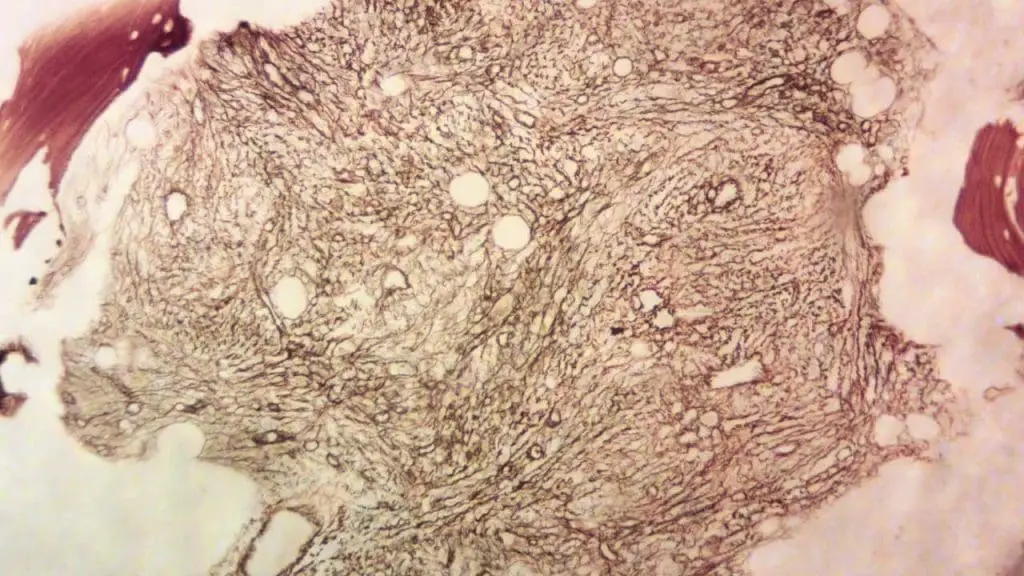
Every immunohistochemistry (IHC) staining run should include positive and negative controls to ensure proper staining and interpretation.
- Positive control: Tissue sections known to express the target antigen, confirming the antibody's functionality and staining effectiveness.
- Negative control: Tissue sections devoid of the target antigen or treated with non-immune serum instead of the primary antibody, revealing background staining levels and non-specific binding.
Tissue Specificity
It's crucial to consider the tissue type and background staining patterns, as some antigens might show inherent expression in specific cell types within the tissue. Analyzing immunohistochemistry (IHC) staining patterns in relation to cell morphology and known tissue architecture helps distinguish specific antibody binding from background noise.
Common IHC Panels in Hematology
In diagnostic hematopathology, we rarely run a single antibody. Instead, we use Panels. Panels are groups of antibodies that work together to confirm a lineage (Myeloid vs. Lymphoid) or sub-classify a disease (e.g., Diffuse Large B-cell Lymphoma vs. Follicular Lymphoma).
The Screening Panel
Clinical Scenario: A lymph node is enlarged, and the pathologist sees abnormal cells. The first step is to determine if it's a lymphoma, a carcinoma (metastatic cancer), or a reactive process.
| Marker | Target | Significance |
| CD45 (LCA) | Leukocyte Common Antigen | Positive in almost all hematologic malignancies (Lymphomas/Leukemias). Negative in carcinomas. |
| Pan-Cytokeratin | Epithelial Cells | Positive in Carcinomas. Negative in Lymphomas. |
| S100 / SOX10 | Melanocytes | Positive in Melanoma (which can mimic lymphoma). |
| CD3 | T-cells | Identifies T-cell lineage. |
| CD20 | B-cells | Identifies B-cell lineage. |
The Small B-Cell Lymphoma Panel
Clinical Scenario: The patient has low-grade lymphocytosis or enlarged nodes with small, monotonous lymphocytes. We need to distinguish between CLL/SLL, Mantle Cell Lymphoma, and Follicular Lymphoma.
Core Markers: CD5, CD10, CD23, Cyclin D1, BCL-2, BCL-6.
Decision Tree
- CD5 Positive?
- YES: It's likely CLL/SLL or Mantle Cell Lymphoma.
- Next Step: Check CD23 and Cyclin D1.
- CLL/SLL: CD23 (+), Cyclin D1 (-).
- Mantle Cell: CD23 (-), Cyclin D1 (+).
- NO: Check CD10.
- YES: It's likely CLL/SLL or Mantle Cell Lymphoma.
- CD10 Positive?
- YES: Likely Follicular Lymphoma or Burkitt Lymphoma.
- Distinguisher: BCL-2. Follicular is BCL-2 (+); Burkitt is BCL-2 (-).
- NO: Likely Marginal Zone Lymphoma or LPL (diagnosis of exclusion).
- YES: Likely Follicular Lymphoma or Burkitt Lymphoma.
The Large Cell Panel (Aggressive Lymphomas)
Clinical Scenario: Large, ugly cells are destroying the lymph node architecture. The top differential is Diffuse Large B-Cell Lymphoma (DLBCL) vs. Hodgkin Lymphoma.
| Marker | Hodgkin Lymphoma (cHL) | DLBCL |
| CD45 | Usually Negative | Positive |
| CD20 | Variable (often weak/neg) | Strongly Positive |
| CD30 | Strongly Positive | Variable (can be +) |
| CD15 | Positive (in ~75% of cases) | Negative |
| PAX5 | Weak (Dim) Nuclear + | Strong Nuclear + |
| EBV (LMP1) | Positive in ~40% cases | Variable |
Hans Algorithm (for DLBCL Subtyping): Once DLBCL is confirmed, we use CD10, BCL-6, and MUM1 to classify it as "Germinal Center B-cell (GCB)" type (better prognosis) or "Non-GCB" type.
The Acute Leukemia Panel (Blast Identification)
Clinical Scenario: A bone marrow biopsy is packed with immature "blasts." We need to know if it is AML (Myeloid) or ALL (Lymphoid).
- General Blast Marker: CD34 (Note: Not all leukemias are CD34+).
- Myeloid Markers (AML):
- MPO (Myeloperoxidase): The most specific marker for myeloid lineage.
- CD117 (c-Kit): Marker for immature myeloid cells/blasts.
- CD13 / CD33: Pan-myeloid markers.
- CD68 / CD163: Monocytic differentiation (for Acute Monoblastic Leukemia).
- Lymphoid Markers (ALL):
Plasma Cell Neoplasm Panel (Multiple Myeloma)
Clinical Scenario: High serum protein or lytic bone lesions. We are looking for clonality in plasma cells.
- CD138 (Syndecan-1): The "Gold Standard" plasma cell marker. Stains the membrane intensely.
- CD38: Also stains plasma cells but can stain other cells too.
- Kappa & Lambda Light Chains:
- Normal plasma cells are a mix of Kappa and Lambda.
- Myeloma cells are Restricted (e.g., all Kappa or all Lambda).
- Aberrant Markers:
- CD56: Normal plasma cells are CD56(-). Myeloma cells are often CD56(+).
- Cyclin D1: Positive in ~30% of Myeloma (t(11;14) translocation).
Common "Look-Alike" Differentiations
| If you are trying to separate... | Use this Marker |
| CLL vs. Mantle Cell | Cyclin D1 (Mantle is +, CLL is -) |
| Follicular vs. Reactive Hyperplasia | BCL-2 (Follicular is +, Reactive is -) |
| Hodgkin vs. Anaplastic Large Cell (ALCL) | ALK-1 (ALCL is often +, Hodgkin is -) |
| Seminoma vs. Lymphoma | OCT3/4 or SALL4 (Seminoma is +) |
Common IHC Panels by Lymphoma Subtype
| Lymphoma Category | Subtype | Key IHC Markers (Positive) | Key IHC Markers (Negative) | Clinical "Key" |
| Small B-Cell | CLL / SLL | CD5, CD23, CD20 (weak), LEF1 | Cyclin D1, CD10 | LEF1 is the most specific nuclear marker for CLL. |
| Mantle Cell (MCL) | CD5, CD20, Cyclin D1, SOX11 | CD23, CD10 | t(11;14) drives the Cyclin D1 expression. | |
| Follicular (FL) | CD10, BCL2, BCL6, CD20 | CD5, CD23 | BCL2+ follicles distinguish it from reactive centers. | |
| Marginal Zone (MZL) | CD20, BCL2, MNDA | CD5, CD10, CD23 | A "diagnosis of exclusion" in small B-cell panels. | |
| Large B-Cell | DLBCL | CD20, PAX5, BCL6 (+/-) | CD3 (except reactive) | Use Hans Algorithm (CD10, BCL6, MUM1) for subtyping. |
| Burkitt Lymphoma | CD10, BCL6, Ki-67 (~100%), MYC | BCL2 | The "starry sky" morphology correlates with 100% Ki-67. | |
| T-Cell | PTCL, NOS | CD3, CD2, CD5, CD4 (>CD8) | CD20 | Often shows "loss" of one or more pan-T markers (e.g., CD7-). |
| ALCL | CD30 (strong), ALK (+/-), EMA | CD3 (often negative) | "Hallmark cells" are always strongly CD30 positive. | |
| NK/T-cell (Nasal) | CD3 (cyto), CD56, EBER-ISH, TIA-1 | CD20 | EBV (EBER) is mandatory for this diagnosis. | |
| Hodgkin | Classical HL | CD30, CD15, PAX5 (weak/dim) | CD45, CD3, CD20 (usually) | RS cells are typically CD45 negative. |
| NLPHL | CD20, CD45, OCT2, BOB.1 | CD30, CD15 | "Popcorn cells" stain like mature B-cells, unlike classical HL. | |
| Plasma Cell | Myeloma | CD138, CD38, CD56, Kappa/Lambda | CD20, CD45 | Look for light chain restriction (Kappa:Lambda ratio >3:1). |
The "Big 4" Small B-Cell Differentiator
When a lab receives a "small cell" biopsy, they usually run this four-marker panel immediately to narrow the field.
| Marker | CLL / SLL | Mantle Cell | Follicular | Marginal Zone |
| CD5 | + | + | - | - |
| CD10 | - | - | + | - |
| CD23 | + | - | - | - |
| Cyclin D1 | - | + | - | - |
Specialized "Niche" Markers
If the primary panel is ambiguous, pathologists use these "tie-breakers":
- SOX11: Highly specific for Cyclin D1-negative Mantle Cell Lymphoma.
- TdT: The absolute marker for Lymphoblastic lymphoma/leukemia (immature cells).
- Annexin A1 / BRAF V600E: Used specifically to confirm Hairy Cell Leukemia.
- BCL-2: In Follicular Lymphoma, BCL-2 staining inside the follicles is the "smoking gun" (normal reactive follicles are BCL-2 negative).
Troubleshooting
Remember that immunohistochemistry (IHC) staining intensity and quality can be influenced by technical factors like fixation, tissue processing, and staining protocol variations. Consistent optimization and standardization of procedures are essential for reliable and reproducible results.
The Golden Rule: Check Your Controls First
Before blaming the antibody, look at the slides:
- Positive Control: If this is negative, the reagent or protocol failed.
- Positive Control (Good) + Patient (Negative): The patient truly does not express the antigen (or the tissue is over-fixed).
- Negative Control: If this is brown, you have "background noise" and cannot trust any positive result on the patient slide.
Scenario A: No Staining at All (The "Blank" Slide)
The entire slide is blue (hematoxylin only), with no brown chromogen visible.
| Possible Cause | Solution |
| Missed Step | Did you forget to add the Primary Antibody? (It happens!) Check your pipetting order. |
| Wrong Secondary Antibody | Check the species. Did you use an "Anti-Mouse" secondary on a "Rabbit" primary antibody? They must match. |
| Sodium Azide Interference | Sodium azide (a preservative) destroys HRP (Horseradish Peroxidase). Ensure your buffers are azide-free if using HRP. |
| DAB Issues | Did the DAB mixture turn brown before you put it on the slide? If so, it oxidized in the tube. Mix fresh DAB immediately before use. |
| Antigen Retrieval Failure | The pH of the retrieval buffer might be wrong. EDTA (pH 9.0) is often stronger than Citrate (pH 6.0). Try switching buffers. |
Scenario B: High Background (The "Dirty" Slide)
The whole tissue is covered in a brown haze, making it hard to see specific cells.
| Possible Cause | Solution |
| Endogenous Peroxidase | Crucial for Heme: Red blood cells and granulocytes are packed with peroxidase. Ensure you block with 3% Hydrogen Peroxide (H₂O₂) for 10–15 mins before adding antibodies. |
| Inadequate Blocking | The secondary antibody is sticking to tissue proteins non-specifically. Use a Protein Block (BSA or Serum from the secondary antibody host) for 30 mins before the primary. |
| Primary Too Concentrated | If the antibody is too strong, it binds everywhere. Titrate the antibody (try 1:100, 1:200, 1:500) to find the "sweet spot." |
| Dried Out Tissue | If the slide dries out at any point during the run, the edges will turn dark brown (Edge Effect). Keep slides in a humidity chamber. |
| Dewaxing Issues | Residual paraffin repels water and traps reagents. Change your Xylene and Alcohols frequently. |
Scenario C: Weak or Patchy Staining
The signal is there, but it is too faint to be diagnostic.
| Possible Cause | Solution |
| Over-Fixation | Formalin cross-links proteins. If the tissue sat in formalin for >48 hours (common over weekends), you need more aggressive Antigen Retrieval (longer heat or higher pH). |
| Bone Marrow Decalcification | Hematology Alert: Strong acid decalcifiers (Nitric Acid/RDO) destroy antigens (especially CD4, CD8, and CD30). Use EDTA decalcification for IHC markers, even though it is slower. |
| Old Reagents | Diluted antibodies are unstable. Store concentrated antibodies at 4°C (or -20°C if glycerol-based) and dilute only what you need for the day. |
| Short Incubation | Extend the primary antibody incubation time (e.g., from 1 hour at Room Temp to Overnight at 4°C). |
Specific Tips for Hematology Specimens
1. The "Crush Artifact" in Lymph Nodes
- Problem: Lymph nodes are fragile. If the surgeon or technician squeezes the tissue with forceps, the nuclei smear.
- Result: It looks like DNA dust. IHC markers (like Ki-67 or BCL-6) will stain the smeared DNA, creating false positives or unreadable slides.
- Fix: Handle lymph nodes gently. On the slide, look for areas with intact cellular outlines.
2. Bone Marrow Biopsies (The "Decal" Problem)
- Problem: As mentioned above, acids destroy DNA and proteins. A bone marrow biopsy decalcified in strong acid often yields false negative results for delicate nuclear markers (like TdT, PAX5, and Cyclin D1).
- Fix: If you suspect over-decalcification, try using a Polyclonal antibody (which targets multiple epitopes) instead of a Monoclonal one, or rely on Flow Cytometry for confirmation.
3. Edge Artifacts
- Problem: The edges of a needle core biopsy often stain positively for everything (false positive).
- Reason: Mechanical damage during the biopsy procedure and drying during processing.
- Fix: Ignore the edges. Read the cells in the center of the core.
Frequently Asked Questions (FAQs)
What are the two types of immunohistochemistry (IHC)?
he two main types of Immunohistochemistry (IHC) are:
- Chromogenic IHC: This method uses a labeled antibody that, when bound to its target antigen, produces a visible color change. This allows for direct observation of the antigen's location under a light microscope.
- Fluorescent IHC: In this technique, the antibody is labeled with a fluorescent marker. When the antibody binds to its target, it emits a fluorescent signal that can be detected using a fluorescence microscope. This method is particularly useful for visualizing multiple antigens simultaneously.
What are the most common IHC stains?
Here are some of the most common IHC stains:
- DAB (Diaminobenzidine): A chromogen that produces a brown color when reacted with hydrogen peroxide, often used in conjunction with peroxidase-labeled antibodies.
- AEC (3-amino-9-ethylcarbazole): A chromogen that produces a red color, often used for double staining with DAB.
- Alkaline phosphatase-conjugated antibodies: These antibodies can be detected using substrates like BCIP/NBT (5-bromo-4-chloro-3-indolyl phosphate/nitroblue tetrazolium), which produce a blue or purple color.
- Fluorescent dyes: Examples include FITC (fluorescein isothiocyanate), rhodamine, and Cy3/Cy5. These dyes emit light of different colors when excited by specific wavelengths of light.
Some commonly stained antigens include:
- Ki-67: A marker for cell proliferation.
- CD3: A marker for T lymphocytes.
- CD20: A marker for B lymphocytes.
- ER (estrogen receptor): A marker for breast cancer.
- PR (progesterone receptor): A marker for breast cancer.
- HER2: A marker for breast cancer.
- CK (cytokeratin): A marker for epithelial cells.
- Vimentin: A marker for mesenchymal cells.
- S-100: A marker for neural cells.
These are just a few examples, and the choice of stain depends on the specific research question or clinical application.
What are the three most common problem areas in IHC staining?
Three common problem areas in IHC staining are:
- Background staining: This occurs when the antibody binds nonspecifically to other proteins or components in the tissue. It can interfere with the visualization of the target antigen and make it difficult to interpret the results.
- Weak or inconsistent staining: This can be caused by various factors, such as low antibody concentration, inadequate antigen retrieval, or suboptimal staining conditions. Weak staining may make it difficult to detect the target antigen, while inconsistent staining can lead to unreliable results.
- Endogenous peroxidase activity: In chromogenic IHC using peroxidase-labeled antibodies, endogenous peroxidase activity in the tissue can interfere with the detection of the target antigen. This can lead to false positive staining.
To address these problems, researchers can take several steps, including:
- Optimizing antigen retrieval: This process involves treating the tissue with heat or chemicals to expose the target antigen.
- Using appropriate blocking buffers: Blocking buffers can help to reduce nonspecific binding of the antibody to background proteins.
- Adjusting antibody concentrations: The concentration of the primary and secondary antibodies can be adjusted to optimize staining intensity and reduce background noise.
- Using appropriate controls: Positive and negative controls can help to validate the staining and identify potential problems.
- Considering the tissue type and fixation method: The choice of tissue type and fixation method can influence the staining quality.
- Using endogenous peroxidase blocking reagents: These reagents can inhibit endogenous peroxidase activity and prevent false positive staining.
By carefully addressing these problem areas, researchers can improve the quality and reliability of their IHC staining results.
Why do I need to perform Antigen Retrieval?
Most tissue samples are fixed in formalin, which preserves tissue by creating cross-links between proteins. These cross-links can "mask" or hide the antigen, preventing the antibody from binding. Antigen retrieval breaks these bonds, exposing the antigen for detection.
What is the difference between Direct and Indirect IHC?
In Direct IHC, the primary antibody is labeled with a tag (enzyme/dye) and binds directly to the antigen. It is faster but less sensitive. In Indirect IHC (most common), an unlabeled primary antibody binds to the antigen, and then a labeled secondary antibody binds to the primary. This amplifies the signal because multiple secondary antibodies can bind to a single primary antibody.
Why is my IHC negative control showing positive staining?
This indicates "non-specific background staining." Common causes include:
- Inadequate blocking of endogenous peroxidases or proteins.
- Primary or secondary antibody concentration is too high.
- Tissue drying out during the procedure.
- Inadequate washing between steps.
Can I use IHC on frozen tissue sections?
Yes. Frozen sections (cryosections) do not require deparaffinization or vigorous antigen retrieval because the tissue hasn't been fixed in formalin. However, the morphology is often poorer compared to formalin-fixed paraffin-embedded (FFPE) tissue.
How long can I store cut slides before staining?
It is best to stain slides immediately. However, cut slides can be stored at 4°C for several weeks. Over time (months), antigens can degrade (oxidize), leading to weak signals, especially for nuclear antigens like Ki-67 or p53. Dip-coating slides in paraffin can extend storage time.
Glossary of Related Medical Terms
- Antibody (Immunoglobulin): A Y-shaped protein produced by the immune system that specifically identifies and neutralizes foreign objects like bacteria or viruses. In IHC, it is used to detect specific antigens.
- Antigen: Any substance (usually a protein) that causes the immune system to produce antibodies against it. In IHC, this is the "target" we are trying to visualize in the tissue.
- Antigen Retrieval (Epitope Retrieval): A process, usually involving heat and a specific buffer, used to "unmask" antigens that were cross-linked and hidden during tissue fixation (formalin).
- Chromogen: A chemical substrate (like DAB) that reacts with an enzyme (like HRP) to produce a colored precipitate, making the antibody-antigen complex visible under a microscope.
- Counterstain: A stain (usually Hematoxylin) applied after the primary reaction to make the background tissue visible, providing contrast so the pathologist can see the cell structure surrounding the positive stain.
- Deparaffinization: The removal of paraffin wax from tissue sections using solvents like xylene. This is necessary because antibodies cannot penetrate wax.
- Epitope: The specific part of an antigen that an antibody attaches to.
- HRP (Horseradish Peroxidase): An enzyme commonly conjugated (attached) to secondary antibodies. It catalyzes the reaction that turns the chromogen into a visible color.
- Primary Antibody: The first antibody applied in the protocol; it specifically binds to the target antigen in the tissue.
- Secondary Antibody: An antibody that binds to the primary antibody. It usually carries the visual marker (enzyme or fluorophore) to amplify the signal.
Disclaimer: This protocol is for educational purposes only. Local laboratory standard operating procedures take precedence. It is not intended to be a substitute for informed professional medical advice, diagnosis, or treatment. Always consult a qualified healthcare professional for clinical decision-making. While the information presented here is derived from credible medical sources and is believed to be accurate and up-to-date, it is not guaranteed to be complete or error-free. See additional information.
References
- Kim SW, Roh J, Park CS. Immunohistochemistry for Pathologists: Protocols, Pitfalls, and Tips. J Pathol Transl Med. 2016 Nov;50(6):411-418. doi: 10.4132/jptm.2016.08.08. Epub 2016 Oct 13. PMID: 27809448; PMCID: PMC5122731.
- Bain BJ, Bates I, Laffan MA. Dacie and Lewis Practical Haematology: Expert Consult: Online and Print 12th Edition (Elsevier). 2016.
- Steele J. Histology, Immunohistochemistry and In Situ Hybridisation, Lab Protocols. 2017.
- Nguyen T. Immunohistochemistry: A Technical Guide to Current Practices (Cambridge University Press). 2022.
- Renshaw S. Immunohistochemistry and Immunocytochemistry: Essential Methods (Wiley-Blackwell). 2017.
- Buchwalow IB, Böcker W. Immunohistochemistry: Basics and Methods (Springer). 2010.