Key Takeaways
Cell surface markers are indispensable in hematology, acting as unique “fingerprints” that allow for the precise identification, classification, and functional understanding of various blood cell types. For example, CD34 is a pivotal marker for hematopoietic stem and progenitor cells, crucial not only for stem cell transplantation but also as a prognostic indicator and for distinguishing different acute leukemia subtypes.
| Cell Type | Classical Marker Panel |
| Hematopoietic Stem Cell (HSC) | Lin-CD34+CD38-CD45RA-CD90+CD49f+ |
| Common Lymphoid Progenitor (CLP) | Lin-CD34+CD38-/loCD45RA+CD90- |
| Common Myeloid Progenitor (CMP) | CD34+CD38+CD123medCD135+CD45RA- |
| Erythrocyte (Red Blood Cells) | CD235a (Glycophorin A) |
| T cell | CD3 and CD4 or CD8 |
| B cell | CD19 and CD20 |
| Platelets | CD41 and CD61 |
Cell Surface Markers
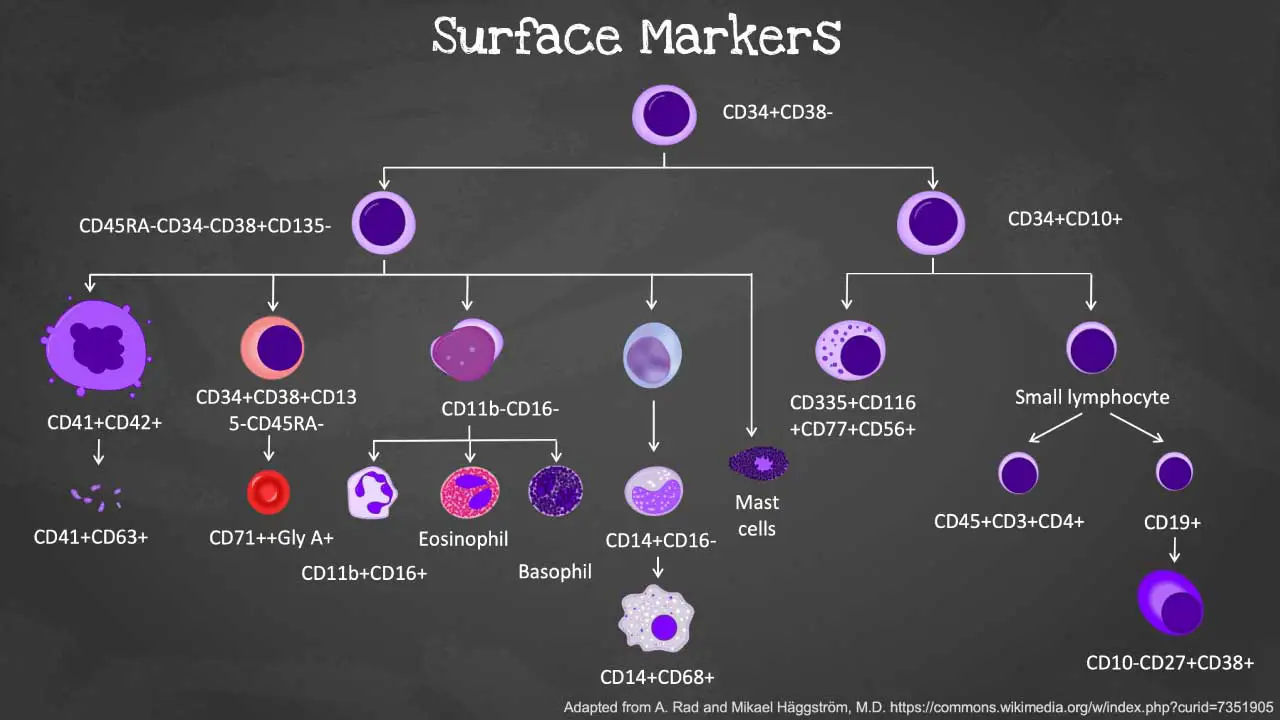
Cell surface markers are protein, carbohydrate, or lipid molecules embedded in the cell membrane. Each type of cell has a specific set of surface markers that makes it unique. Surface markers include molecules that are able to bind various other molecules or cells, or antibodies directed against them. Antibodies recognizing similar antigens are grouped into clusters, and are known as clusters of differentiation (CD). Key CD markers in hematology include CD34, CD38, CD19, CD235a and CD45.
Function of Cell Surface Markers
Cell Type Identification and Classification
Think of cell surface markers like fingerprints. Each cell type possesses a distinct fingerprint of cell surface markers, allowing their identification and classification. This is crucial for:
- Immune System: B cells use specific cell surface markers on pathogens to identify and target them for destruction. T cells recognize infected cells by their altered cell surface markers, triggering an immune response.
- Tissue Development: Different cell types in an organ express specific cell surface markers, guiding their spatial organization and function. For example, markers on brain cells guide their connections to form intricate neural networks.
- Cancer Diagnosis: Tumor cells often display unique cell surface markers not found in healthy cells. These cell surface markers serve as diagnostic tools and potential targets for cancer therapy.
Cell-Cell Interactions and Signaling
Cell surface markers are like handshake points for cellular communication. They act as docking stations for proteins, triggering a cascade of signals within and between cells.
- Cell Adhesion: Specific cell surface markers on one cell bind to complementary ones on another, facilitating adhesion and forming intricate cell-cell structures like tissues and organs.
- Signal Transduction: Ligand-receptor binding between cell surface markers on different cells can activate signaling pathways, leading to diverse cellular responses like growth, differentiation, or migration.
- Immune Regulation: T cells recognize specific cell surface markers on antigen-presenting cells, stimulating the immune response, while other markers on immune cells can dampen inflammation.
Cell Adhesion and Migration
Cell surface markers act like guideposts and anchors for cell movement.
- Directional Migration: Specific cell surface markers interact with molecules in the extracellular matrix, directing their migration towards specific destinations during development or wound healing.
- Cell Adhesion and Detachment: Dynamic changes in cell surface markers’ expression can regulate cell adhesion strength, allowing cells to detach and migrate freely during processes like embryonic development or immune cell trafficking.
- Extracellular Matrix Interaction: Cell surface markers bind to components of the extracellular matrix, providing structural support and influencing cell behavior.
Differential Regulation of Cellular Functions
Cell surface markers act like switches, controlling specific functions within individual cells.
- Cell Activation and Differentiation: Specific cell surface markers on stem cells can be targeted by signaling molecules, triggering their differentiation into specialized cell types.
- Enzyme Regulation: Certain cell surface markers directly regulate the activity of enzymes within cells, influencing metabolic pathways and cellular functions.
- Drug Targeting: Cell surface markers on diseased cells can be targeted by therapeutic drugs, specifically inhibiting their growth or function.
Flow Cytometry for Cell Surface Marker Detection
Flow cytometry is a powerful technique that allows scientists to analyze and sort cells based on their unique patterns of protein expression.
The Principle
Flow cytometry works like a sophisticated sorting machine. Cells suspended in a fluid stream pass through a laser beam one by one. Specific antibodies, each conjugated to a unique fluorescent dye, bind to the target surface markers on the cell.
As the cell passes through the laser, the dye gets excited and emits a characteristic fluorescent signal. This signal is then detected by sensors, revealing the presence and quantity of the targeted surface marker.
Multiparametric Immunophenotyping
Flow cytometry’s true power lies in its ability to analyze multiple markers simultaneously. By using a cocktail of antibodies, researchers can create a detailed profile of a cell’s surface marker expression, like a multi-colored fingerprint. This technique, called multiparametric immunophenotyping, allows scientists to:
- Distinguish different cell types: Unique combinations of cell surface markers define cell identities. For example, identifying the presence of CD3 and CD8 on a cell indicates it’s a cytotoxic T lymphocyte.
- Track cell differentiation: As cells mature, their cell surface marker profile changes. Flow cytometry can track these changes, providing insights into the developmental journey of a cell.
- Study cellular function: Certain cell surface markers are associated with specific cellular functions. Flow cytometry helps correlate marker expression with cellular activity.
Fluorescence-Activated Cell Sorting (FACS)
Flow cytometry isn’t just about observation; it can also be used to isolate specific cell populations. By combining the analysis with cell sorting capabilities, FACS allows researchers to:
- Purify rare cell populations: For studying specific cell types like HSCs, FACS can isolate them from a complex mixture of blood cells.
- Separate cells based on functional characteristics: Cell surface markers associated with specific cellular functions can be used to isolate active or resting cells, for example.
Antibodies: The Key Players
The accuracy and specificity of flow cytometry rely heavily on high-quality antibodies. These molecules are designed to bind to specific cell surface markers with high affinity. By conjugating them to different fluorescent dyes, scientists can create a diverse toolbox for targeted marker detection.
Beyond Flow Cytometry
While flow cytometry reigns supreme, other techniques offer complementary insights.
- Immunohistochemistry: This technique uses antibodies to visualize the localization of cell surface markers within tissue sections.
- Fluorescence microscopy: This technique allows researchers to observe the spatial distribution of cell surface markers on individual cells with high resolution.
Clinical Applications of Cell Surface Markers
The development of more fluorochromes and discovery of cell surface markers allow hematopoietic stem and progenitor cells (HSPCs) to be identified prior to pre-transplant in a research setting and later the engraftment of target cell populations are possible to be analyzed in a single test.
Also, hematopoietic cells either at its quiescent or activated state can be identified and the aberrancies of blood can be diagnosed.
Diagnosis and Classification
- Leukemias: Different leukemia types express distinct cell surface markers, allowing classification of the disease and proper treatment accordingly. For example, B-cell acute lymphoblastic leukemia (ALL) expresses CD19 and CD38, while T-cell ALL lacks these markers and expresses CD7.
- Lymphomas: Cell surface markers like CD20 on B cells and CD3 on T cells help distinguish different lymphoma subtypes, guiding treatment strategies.
- Anemias: Aplastic anemia exhibits decreased expression of human leukocyte antigen (HLA) markers on bone marrow cells.
Monitoring Disease Progression and Treatment Response
- Monitoring minimal residual disease (MRD) in leukemia patients after therapy using specific cell surface marker detection can predict relapse risk and guide treatment adjustments.
- Measuring cell surface marker expression on immune cells can predict response to immunotherapy in cancer patients.
Identification and Isolation of Specific Cell Populations
- Hematopoietic stem cell transplantation for leukemia and other blood cell disorders relies on identifying and isolating HSCs based on specific cell surface markers like CD34 and CD133.
- CAR T-cell therapy for cancer involves engineering T cells with receptors targeting specific cell surface markers on cancer cells.
Development of Targeted Therapies
- Monoclonal antibodies targeting specific cell surface markers on cancer cells can deliver targeted drugs or toxins directly to the tumor.
- Small-molecule drugs can be designed to interfere with signaling pathways activated by specific cell surface markers.
Introduction to Hematopoiesis
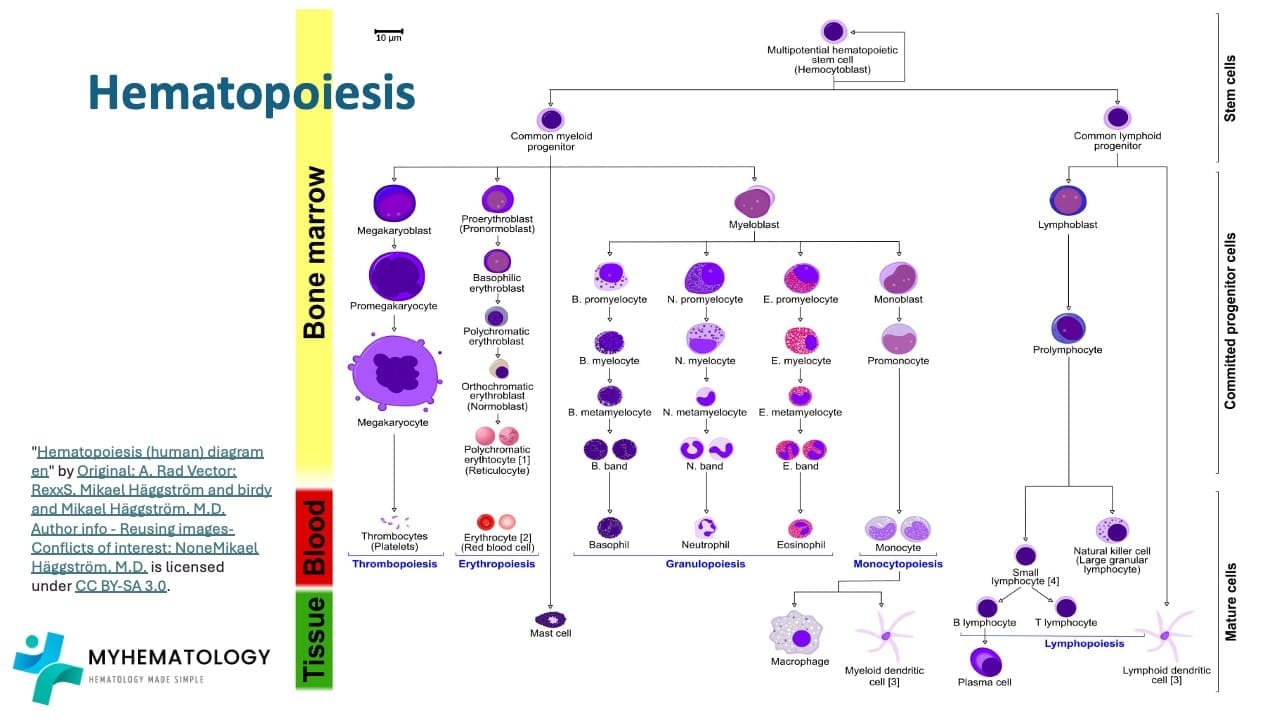
Hematopoiesis is the remarkable process by which the body creates all the different types of blood cells including the red blood cells, white blood cells and platelets.
During this process of blood production, the different blood cells are derived from the hematopoietic stem and progenitor cells (HSPC) through a tightly coordinated sequence of events.
The mother cell from which all blood cells originate from is the hematopoietic stem cell (HSC). HSCs have the ability to self-renew, which means they can divide to produce more stem cells and they can differentiate into a limited number of cell types within a specific lineage (multipotent).
From the HSCs, hematopoietic progenitor cells (HPC) such as the common lymphoid progenitors (CLP), granulocyte-macrophage progenitors (GMP) and megakaryocyte-erythroid progenitors (MEP) arises to give differentiate into hematopoietic precursors, then become increasingly lineage-specific and ultimately differentiate into all lineages of mature hematopoietic cells.
The CLP will differentiate further to develop into T-, B- and natural killer (NK-) cells. These cells are immune cells that are a well-coordinated army defending our body against invaders.
The GMPs form granulocyte-macrophage lineage cells such as neutrophils, basophils, eosinophils, mast cells, dendritic cells or monocytes; which are frontline defenders against infections, whereas the megakaryocytes and platelets as well as red blood cells derive from MEPs.
Differentially expressed cell surface markers serve as vital tools for isolating and characterizing HSCs and their lineage-restricted progenitors within the complex hematopoietic hierarchy.
Multifaceted Roles of CD34
CD34, a transmembrane phosphoglycoprotein, is far more than just a cell surface marker for hematopoietic stem and progenitor cells (HSPCs); its multifaceted roles extend across various cell types and clinical applications, making it a pivotal molecule in biological research and medicine.
Beyond its well-known presence on HSPCs, CD34 expression is also observed on non-hematopoietic cells, including interstitial cells, endothelial cells, fibrocytes, and muscle satellite cells.
In muscle satellite cells, also known as muscle stem cells, CD34 serves as a key marker, playing a fundamental role in regulating muscle progenitor cell differentiation and maintaining the satellite cell population.
Its presence on endothelial cells is significant for vascular development and integrity. Furthermore, CD34 expression has been identified on various cancer stem cells, highlighting its emerging importance in cancer research.
Function of CD34
Regarding its CD34 function, evidence strongly suggests that its primary role is in cell adhesion, rather than directly in proliferation signaling. Studies have shown that the cytoplasmic domain of CD34 is essential for cytoadhesion signaling, mediating the attachment of stem cells to the bone marrow extracellular matrix or stromal cells.
In contrast, it does not appear to contain the necessary elements to transduce a proliferative signal in hematopoietic cells, distinguishing its adhesive function from growth-promoting pathways.
A significant shift in understanding has occurred with the “lineage priming” hypothesis, which posits that HSCs promiscuously express lineage-specific genes before commitment. This has been confirmed by findings that
CD34-positive HSCs can express myeloid markers such as CD13, CD33, and CD123. This discovery overturns the previous dogma that human HSCs are devoid of lineage-specific antigens and carries profound
Clinical Significance of CD34
CD34 expression in leukemia holds significant prognostic value and aids in diagnosis and classification. It is widely used to identify blast cells in acute leukemias of any lineage.
In conjunction with other markers like CD10, CD19, CD5, and CD56, CD34 helps distinguish between various acute leukemias (B-cell ALL, T-cell ALL, and acute myeloid leukemia), myelodysplastic syndromes (MDS), and myeloproliferative neoplasms (MPN). Furthermore, CD34, alongside CD38, serves as a prognostic biomarker for acute B lymphoblastic leukemia (B-ALL), where the absence of CD34 or high CD38 expression can correlate with a favorable prognosis.
This makes CD34 an indispensable tool for both diagnostic precision and predicting patient outcomes in hematological malignancies.
Beyond its roles in stem cell biology and leukemia, CD34 and angiogenesis are closely linked. CD34-positive cells, particularly those derived from bone marrow, have demonstrated utility in therapeutic angiogenesis for cardiovascular diseases. These cells can induce the formation of new blood vessels in ischemic tissues, offering a novel therapeutic strategy for conditions like myocardial, peripheral, and cerebral ischemia.
CD34: Various Roles & Clinical Significance
| Context/Cell Type | Key Role/Function of CD34 | Clinical/Biological Implication of CD34 |
| Hematopoietic Stem/Progenitor Cells | Stem cell identification & enrichment Adhesion | Bone marrow transplantation Cell sorting |
| Muscle Satellite Cells | Muscle stem cell marker Regulation of differentiation | Muscle regeneration Myogenesis |
| Endothelial Cells | Cell-cell adhesion Angiogenesis | Therapeutic angiogenesis in CVD Vascular development |
| Leukemia Diagnosis/Prognosis | Blast cell identification Prognostic biomarker | Classifying leukemias (AML, ALL) Predicting outcomes |
| Cancer Stem Cells | Marker for various cancer stem cell populations | Research into cancer initiation and recurrence |
| Expression of Myeloid Markers on HSCs | Lineage priming Promiscuous gene expression | Implications for targeted AML therapies HSC killing |
Differential Cell Surface Markers of Hematopoietic or Blood Cells
| Cell type | Cell Surface Markers | References |
| Hematopoietic stem cells (HSCs) | CD34+CD38- -Lin−CD34+CD38−CD90+CD45RA− Lin−CD34+CD38−CD49f+CD45RA−CD90+/− Lin-CD34+CD38-CD45RA-CD90+CD49f+ | Classical Majeti et al., 2007 Wang, 2012 Qiao et al., 2014 Weeda et al., 2022 |
| Common lymphoid progenitors | CD34+CD10+ Lin-CD34+CD38-/loCD45RA+CD90- Lin−CD34+CD10+ (bone marrow) CD34+CD38-CD45RA+CD7+ (cord blood) | Classical Hokland et al., 1987 Hao et al., 2001 |
| Common myeloid progenitors | CD45RA-CD34+CD19-IL-3R⍺low/-CD110- Lin−CD34+CD38+CD135+CD45RA−CD7−CD10− CD45RA-CD34-CD38+CD135- (BM and cord blood) CD34+CD38+CD123intCD135+CD45RA- | Edvardsson et al., 2006 Qiao et al., 2014 Wang, 2012 Weeda et al., 2022 |
| Granulocyte-macrophage progenitors | CD34+CD123+ CD19-CD34+IL-3R⍺low/-CD45RA+CD110- | Huang et al., 1999 Edvardsson et al., 2006 Weeda et al., 2022 |
| Megakaryocyte-erythroid progenitors | CD34+CD123- CD19-CD34+IL-3Ralow/-CD45RA-CD110+ Lin−CD34+CD38+CD135−CD45RA−CD7−CD10− CD34+CD38+CD135-CD45RA- | Huang et al., 1999 Edvardsson et al., 2006 Qiao et al., 2014 Wang, 2012 Weeda et al., 2022 |
| T cells | CD45+CD3+CD4+ CD4+ (T helper cells) CD25+CD127+FoxP3+ (regulatory T cells) CD3 and CD4 or CD8 | O’Gorman and Gelman, 1997 Romagnani, 1991 Fontenot et al., 2003; Yu et al., 2012 |
| Natural killer cells | CD16+ (peripheral blood) CD56+ (mucosal) CD56+ or CD16−CD56− (lymph nodes) | Mandelboim et al., 1999 Reeves et al., 2010 Reeves et al., 2010 |
| B cells | CD19+ (classical) CD10+CD27−CD38+ (immature) CD10−CD27−CD38− (naïve) CD10−CD27+CD38− (memory) CD10−CD27+CD38+ (plasma) | Carter et al., 2002 Wang et al., 2012 Baez et al., 2014 Caraux et al., 2010; Pritz et al., 2015 |
| Monocytes | CD14+ (general) CD14+CD16− (classical) CD14++CD16+ (intermediate) CD14+CD16++ (non-classical) | Martin et al., 1994 Zamani et al., 2013 Passlick et al., 1989 Sulicka et al., 2013 |
| Dendritic cells | MHC II+Lin- CD1c+CD141+ (myeloid derived) CD123+CD303+ CD304+ (plasmacytoid-derived) | Villadangos et al., 2001 Jongbloed et al., 2010 |
| Granulocytes | CD45+CD11b+CD16+ (immature) CD11b─CD16─ (promyelocyte) CD11b+CD16─ (metamyelocyte) CD11b+CD16+ (mature neutrophils) CD66b+ (eosinophil) CD13+CD107a+CD164+ (activated basophils) | Fujimoto et al., 2000 Guerin et al., 2014 Mazzone and Ricevuti, 1995 Fujimoto et al., 2000 Daley et al., 2008 Yoon et al., 2007 Hennersdorf et al., 2005 |
| Erythroid cells | CD71++gly A+ | Rogers et al., 1996 |
| Platelets | CD41 and CD61 PAC-1+CD63+ CD41+CD62P+ CD42b | van Velzen et al., 2012 |
Differential Cell Surface Markers of Leukemias
High-Yield Flow Cytometry Markers: AML vs. B-ALL vs. T-ALL
| Leukemia Type | Definitive Lineage Markers (The “Must-Knows”) | Secondary / Maturation Markers | Immaturity / Stem Cell Markers |
|---|---|---|---|
Acute Myeloid Leukemia (AML) | MPO (Myeloperoxidase) CD117 (c-kit) | CD13, CD33 CD15, CD14/CD64 (Monocytic) | CD34 HLA-DR (Note: APL/M3 is typically HLA-DR negative) |
| B-Cell Acute Lymphoblastic Leukemia (B-ALL) | cCD79a (cytoplasmic) Strong CD19 | CD10 (CALLA – Common ALL Antigen) CD22, CD24, CD20 (CD20 appears later in maturation) | TdT (Terminal deoxynucleotidyl transferase) CD34, HLA-DR |
| T-Cell Acute Lymphoblastic Leukemia (T-ALL) | cCD3 (cytoplasmic – highly specific) CD7 (highly sensitive, often the first to appear) | CD5, CD2 CD4, CD8, CD1a (indicates cortical stage) | TdT CD34 (Often negative in T-ALL compared to B-ALL/AML) |
Notes:
- The MPO vs. TdT Rule: Myeloperoxidase (MPO) is the undisputed hallmark of myeloid differentiation, while Terminal deoxynucleotidyl transferase (TdT) is the classic nuclear hallmark of immature lymphoid cells (lymphoblasts).
- Cytoplasmic vs. Surface: For definitive early diagnosis, intracellular (cytoplasmic) markers are often more reliable than surface markers. For example, cCD3 is strictly required to define T-cell lineage, whereas surface CD3 is frequently absent on immature T-lymphoblasts.
- Aberrant Expression is Common: It is relatively common for B-ALL blasts to aberrantly express myeloid markers like CD13 or CD33. This does not automatically make it a Mixed-Phenotype Acute Leukemia (MPAL) unless specific lineage-defining criteria are met.
- The CD10 (CALLA) Factor: CD10 is an excellent marker for “Common” B-ALL, which generally carries a more favorable prognosis in children compared to pro-B ALL (which is CD10 negative).
Specific Leukemia Cell Surface Markers
| Disorders | Cell surface marker characteristics |
| Precursor B-ALL | CD45dim-negativeCD19+CD10+CD34+CD22+CD38+cytoCD79a+TdT+CD9+CD58+Cd33+CD20-kappa-lambda- |
| Philadelphia-like B-ALL | CRLF2dim-intCD45dim-negativeCD19+CD10+CD34dim-int |
| T-ALL | CD45+CD7+CD4+CD8+CD1a+CD5+Cyto CD3+TdT+CD2+sCD3-CD16-CD56-CD34-αβ-TCR-γδ-TCR- |
| Early-T-precursor (ETP) ALL | CD7+CD34+CD5dimpartialCD45dim-negativeCD4-CD8-CD1a- |
| AML-M1 | CD45dimCD34dimCD117+HLA-DR+CD38+CD13+CD33+CD15+CD123+MPO+CD71-CD64-CD14-lyso- |
| AML-M2 | MPO+CD11b+CD13+CD15+CD33+CD65+HLA-DR+CD34+CD38+CD117+ |
| AML-M3 | CD45intCD117+CD38+CD13+CD33+CD15dimCD64dimpartialCD71partialMPO+lysopartialCD34-HLA-DR-CD114-CD11b-CD11c- |
| AML-M5 | CD45+HLA-DR+CD38+CD33+CD123+CD64+CD11b+CD11c+lyso+CD117-CD34-MPO- |
| AML-M6 | CD71+glycoA+CD36+CD117+CD34-HLA-DR- |
| AML-M7 | CD41+CD61+CD42b+CD34-HLA-DR- |
| AML-M7 with RAM phenotype | CD34+CD56+CD117+CD61dimCD38dim-negativeCD45dim-negativeHLA-DR- |
| Blastic plasmacytoid dendritic cell neoplasm | CD4+CD56+CD23+CD303+CD304+TCF4+TCL1+CD45dim-intCD7+CD33+CD3-CD14-CD19-CD34-lyso-MPO- |
| Burkitt leukemia | CD45+CD19+CD10+CD20+CD38+kapparestriction |
| CLL | CD20+CD22+CD23+FMC-7-sIg+(weak with light chain restriction)CD200+LEF1+ |
| Mantle cell lymphoma | CD20+sIg+CD23-or dimFMC-7+CD200-LEF1- |
| Hairy cell leukemia | CD20+CD11c+CD25+CD103+CD22+sIg+ with light chain restriction FMC-7+CD23-CD5-CD10- |
| Marginal zone lymphoma | Lack of combine positivity for CD11c, CD25, CD103Lack of bright staining for CD20, CD22 |
| Plasma cell leukemia | CD38+CD138+CD19-CD20- |
Conclusion
In conclusion, cell surface markers are invaluable tools for understanding and regulating hematopoiesis. As research delves deeper and discovers new markers, they pave the way for personalized medicine and developing innovative therapies for blood-related diseases.
They play an essential role as different blood cell types express unique cell surface marker profiles, allowing researchers to distinguish between HSCs (CD34+CD38-), progenitors, and mature cells. This helps map the intricate developmental pathways within the hematopoietic system.
Specific cell surface markers mediate essential functions like cell adhesion, migration, and signaling, providing insights into the mechanisms governing blood cell development and function. Abnormal expression of certain markers can be used to diagnose and classify blood disorders like leukemias and lymphomas, guiding treatment strategies.
In development of research and innovative therapies, specific cell surface markers can be targeted to isolate and manipulate specific cell populations, such as HSCs (CD34+CD38-) for transplantation or CAR T-cell therapy, drugs can be designed to target specific cell surface markers on diseased cells, offering targeted therapies with fewer side effects and introducing genes encoding specific markers can modify the behavior of blood cells, potentially leading to new treatment options.
Frequently Asked Questions (FAQs)
What are cell surface markers on a blood test?
“Cell surface markers on a blood test,” is not referring to markers literally present on the surface of the blood itself, but rather on the various cells found within the blood.
Specifically, these cell surface markers are proteins, carbohydrates, or lipids located on the outer membranes of white blood cells, which play a crucial role in the immune system.
These unique cell surface markers , also known as clusters of differentiation (CDs), act like identifying badges for different types of white blood cells. By analyzing the specific combination of markers present on these cells in a blood test, we can gain valuable insights into the immune system function and diagnose potential health issues.
Are cell surface markers and CD the same?
No, cell surface markers and CDs are not exactly the same, but they are closely related.
Cell surface markers
These are the broader term, referring to any molecule (protein, carbohydrate, lipid) located on the outer surface of a cell. They can have various functions, like cell-cell communication, adhesion, and immune recognition. There are thousands of different cell surface markers on various cell types throughout the body.
CD (cluster of differentiation)
This is a nomenclature system used specifically for surface markers on leukocytes (white blood cells).
Each CD marker has a unique number assigned to it (e.g., CD4, CD8, CD19, CD34). This system helps standardize the identification and characterization of leukocytes based on their surface marker expression.
While the CD (cluster of differentiation) system was initially established to classify surface markers on leukocytes (white blood cells), its application extends significantly beyond just mature white blood cells to cover a broad range of hematopoietic cells including hematopoietic stem cells which are CD34+CD38-.
What do CD markers mean?
CD markers, short for cluster of differentiation, are a system used to classify and identify specific molecules found on the surface of leukocytes, also known as white blood cells. Each CD marker is assigned a unique number, such as CD34, CD4 or CD8, allowing researchers and healthcare professionals to easily refer to them and understand their functions.
Identification and Characterization
Different blood cell types, like B cells, T cells, and monocytes, express distinct combinations of CD markers for example HSCs are CD34+CD38-. This unique “fingerprint” allows scientists to distinguish different cell types and understand their roles in the immune system.
Function and Significance
Each CD marker has a specific function, such as mediating cell-cell communication, adhesion, or activation. Understanding these functions sheds light on how blood cells work and interact with each other.
Diagnosis and Classification
Abnormal expression of specific CD markers can be indicative of various health conditions, including:
- Infections: Certain marker patterns might suggest bacterial, viral, or parasitic infections.
- Immune system disorders: Autoimmune diseases and immunodeficiencies often show altered CD marker profiles on different immune cell types.
- Blood cancers: Leukemias and lymphomas have characteristic marker expression patterns that aid in their diagnosis and classification.
Monitoring and Prognosis
By tracking changes in CD marker expression over time, we can evaluate the effectiveness of treatment for certain conditions like leukemia or monitor immune cell function after organ transplantation.
Further Research and Applications
Researchers are constantly discovering new CD markers and their functions, contributing to our understanding of the immune system and developing novel diagnostic and therapeutic tools.
Can specific cell surface marker profiles on individual HSCs predict their differentiation potential and therapeutic value in transplantation?
The question of whether specific cell surface marker profiles on individual HSCs (CD34+CD38-) can predict their differentiation potential and therapeutic value in transplantation is a fascinating and actively researched area in hematology. While a definitive answer hasn’t been reached yet, promising developments and ongoing research shed light on the potential.
Current Understanding
- Heterogeneity within HSCs: HSCs, despite appearing alike under a microscope, exhibit significant heterogeneity in their differentiation potential and self-renewal capacity.
- Cell surface markers capture some variation: Studies have identified various surface markers associated with specific differentiation biases (e.g., towards B-cells, T-cells, or myeloid lineages).
- Limitations exist: Existing marker panels (CD34+CD38-) only partially capture the complex differentiation potential of HSCs, and predicting therapeutic value remains challenging.
Promising Directions
- Single-cell analysis: Technologies like single-cell RNA sequencing are revealing deeper insights into individual HSC profiles (CD34+CD38-) , including gene expression and functional differences.
- Novel markers: Identifying new surface markers specific to subpopulations of HSCs with distinct properties could improve prediction accuracy.
- Functional assays: Developing assays that test the differentiation potential or therapeutic efficacy of individual HSCs based on their marker profiles (CD34+CD38-) would be invaluable.
Challenges and Considerations
- Technical limitations: Single-cell analysis techniques are still evolving, and isolating and analyzing individual HSCs without altering their properties (CD34+CD38-) remains a challenge.
- Data interpretation: Integrating vast amounts of data from single-cell analysis with functional assays requires advanced computational and statistical approaches.
- Ethical considerations: Manipulating HSCs based on their predicted properties raises ethical concerns that need careful consideration.
Overall, while predicting individual HSC differentiation and therapeutic value solely based on cell surface markers remains elusive, continued research in single-cell analysis, novel marker identification, and functional assays holds tremendous promise for personalized medicine in stem cell transplantation.
Additional Insights
- Studying epigenetics and metabolic profiles alongside cell surface markers might provide a more comprehensive understanding of HSC potential.
- Integrating information from diverse data sources using machine learning algorithms could improve prediction accuracy.
- Collaboration between researchers, clinicians, and ethicists is crucial to ensure responsible development and application of these technologies.
How do microenvironmental cues influence cell surface marker expression during hematopoiesis and how can this be leveraged for therapeutic targeting?
The journey of a blood cell from a humble hematopoietic stem cell (HSC) to its specific mature form is guided by a complex interplay between intrinsic factors and extrinsic cues within the bone marrow microenvironment (BME).
Among these extrinsic cues, microenvironmental signals play a crucial role in regulating the expression of cell surface markers on HSCs (CD34+CD38-) and their progeny, directing their differentiation, proliferation, and survival.
How do microenvironmental cues influence cell surface marker expression?
Various factors in the bone marrow environment (BME)
- Growth factors: Cytokines like SCF, IL-7, and G-CSF interact with specific receptors on HSCs (CD34+CD38-), influencing their survival, proliferation, and commitment towards specific lineages. This often leads to changes in cell surface marker expression reflecting their changing potential.
- Adhesion molecules: Interactions between HSCs and stromal cells through molecules like VCAM-1 and N-cadherin provide physical contact and critical signals that impact gene expression and, consequently, cell surface marker profiles.
- Metabolic factors: Hypoxia (low oxygen) within the BME promotes self-renewal of HSCs, while increased oxygen levels favor differentiation. This metabolic shift can also influence the expression of certain cell surface markers (CD34+CD38-).
- Niche signals: Specialized niches within the BME provide unique signals to HSCs depending on their location. For example, osteoblastic and endothelial niches offer distinct cues that influence cell surafce marker expression and differentiation potential.
Leveraging this knowledge for therapeutic targeting
Understanding how microenvironmental cues influence cell surface marker expression opens exciting avenues for therapeutic strategies.
- Manipulating BME factors: By introducing specific growth factors or mimicking niche signals, scientists can potentially guide HSCs towards desired differentiation pathways, overcoming limitations in current HSC transplantations.
- Targeting marker-specific pathways: Identifying signaling pathways activated by specific microenvironmental cues that lead to altered marker expression could help develop drugs that modulate these pathways, influencing HSC behavior for therapeutic benefit.
- Designing marker-based therapies: Utilizing specific cell surface markers expressed on HSCs (CD34+CD38-) or their progeny allows for targeted delivery of drugs or gene therapies, enhancing treatment efficacy and reducing side effects.
Challenges and future directions
While promising, this field requires further exploration.
- Complexity of the BME: Deciphering the intricate and dynamic interplay of various factors within the BME remains a challenge.
- Limited understanding of signaling pathways: More research is needed to understand the precise downstream effects of microenvironmental cues on specific surface marker expression and HSC function.
- Translational challenges: Bridging the gap between in vitro studies and clinical applications requires careful evaluation of safety and efficacy in animal models and clinical trials.
Can circulating tumor cells (CTCs) isolated based on their surface marker profiles provide valuable prognostic and therapeutic information in leukemia patients?
The potential of circulating tumor cells (CTCs) isolated based on their cell surface marker profiles for providing valuable prognostic and therapeutic information in leukemia patients is an exciting and actively researched area.
Potential Advantages
- Liquid biopsy: CTCs offer a “liquid biopsy” alternative to invasive bone marrow biopsies, providing minimally invasive access to tumor information.
- Heterogeneity: Analyzing individual CTCs with specific surface marker profiles can capture the heterogeneity within a leukemia and identify subpopulations with distinct behavior and therapeutic vulnerabilities.
- Minimal residual disease (MRD) detection: Sensitive detection of rare CTCs even after treatment completion (MRD) can predict relapse risk and guide treatment adjustments.
- Therapeutic targeting: Identifying unique surface markers on CTCs could pave the way for targeted therapies or personalized medicine approaches.
Current Challenges
- Rarity and heterogeneity: CTCs are extremely rare in blood, and leukemia exhibits significant heterogeneity, making isolation and analysis challenging.
- Limited sensitivity and specificity: Existing technologies might not be able to reliably distinguish leukemic CTCs from healthy cells or other blood-borne tumor cells.
- Functional significance: Understanding the functional significance of specific surface marker profiles on CTCs in the context of leukemic progression and relapse is still evolving.
- Clinical validation: More research and clinical trials are needed to fully validate the prognostic and therapeutic value of CTC analysis in leukemia.
Promising Directions
- Advanced technologies: Single-cell analysis, microfluidic devices, and machine learning algorithms are being developed to improve sensitivity, specificity, and capture of relevant information from CTCs.
- Combination of markers: Utilizing panels of surface markers specific to different leukemia subtypes or disease stages could enhance diagnostic and prognostic accuracy.
- Functional assays: Integrating CTC analysis with functional assays to assess drug sensitivity or resistance could guide personalized treatment strategies.
- Early detection and MRD monitoring: Utilizing CTC analysis for early detection of minimal residual disease could improve treatment outcomes and prevent relapse.
How can we integrate information from cell surface markers with other omics data (e.g., genomics, metabolomics) to develop more comprehensive models of leukemia progression and treatment response?
Integrating information from cell surface markers with other omics data is a powerful approach to gain a more comprehensive understanding of leukemia progression and treatment response. By combining diverse data sets, researchers can create more accurate and informative models that could eventually lead to personalized medicine strategies.
Benefits of integration
- Unveiling complex interactions: Omics data provides insights into different layers of cellular information, from genes and proteins to metabolites. Integrating surface marker data with these can uncover how genetic mutations, metabolic pathways, and protein expression interact, influencing disease progression and drug response.
- Identifying novel targets: By analyzing correlations between cell surface markers and other omics data, researchers can identify potential new therapeutic targets that were not evident from looking at each data set individually.
- Personalized medicine: Integrating information from individual patients’ cell surface markers and other omics data can pave the way for personalized treatment plans tailored to their specific disease characteristics and predicted response to specific drugs.
- Improved risk stratification: Integrating omics data can help better categorize patients into risk groups based on their molecular profiles, allowing for more targeted prevention and treatment strategies.
Challenges and considerations
- Data complexity: Analyzing and integrating massive datasets from multiple omics platforms requires advanced computational tools and expertise.
- Data standardization: Omics data generated from different platforms and labs can vary in format and quality, necessitating standardization for effective integration.
- Biological interpretation: Extracting meaningful biological insights from complex multi-omics data requires robust statistical methods and domain knowledge.
- Ethical considerations: Integrating patient-specific omics data raises ethical concerns regarding privacy, data ownership, and potential discrimination based on genetic information.
Promising approaches
- Machine learning and AI: These tools can help analyze vast omics datasets, identify hidden patterns, and build predictive models of disease progression and drug response.
- Network biology: Constructing networks that integrate different omics data can reveal how various molecules and pathways interact, leading to a deeper understanding of disease mechanisms.
- Single-cell analysis: Analyzing individual cells with multi-omics approaches can capture heterogeneity within tumors and identify rare cell populations with specific properties.
Overall, integrating surface marker data with other omics information holds immense potential for developing more comprehensive models of leukemia and guiding personalized treatment strategies. Addressing the challenges and ethical considerations while utilizing advanced analytical tools and collaborative research efforts will be crucial to unlocking the full potential of this promising approach.
What is the difference between CD34 and CD38 in hematopoiesis?
CD34 is a classic marker of primitive hematopoietic stem and progenitor cells. CD38 is an activation and differentiation marker that appears as cells mature. Therefore, the most primitive, self-renewing stem cells are typically identified by a CD34-positive and CD38-negative (CD34+/CD38-) profile.
How does flow cytometry differentiate between myeloid and lymphoid leukemias?
Flow cytometry utilizes specific panels of fluorescently labeled antibodies to map the surface markers of the abnormal cells. Myeloid leukemias will express markers like CD13, CD33, and Myeloperoxidase (MPO), while lymphoid leukemias will express markers specific to B-cells (e.g., CD19, CD79a) or T-cells (e.g., CD3, CD7).
What does it mean when a cell population is described as “Lineage Negative” (Lin-)?
When a cell population is Lineage Negative, it means the cells do not express the surface markers associated with mature, fully differentiated blood cells (such as CD3 for T-cells, CD19 for B-cells, or CD14 for monocytes). In the lab, researchers use a “lineage cocktail” of antibodies to exclude mature cells so they can isolate the rare, immature stem and progenitor cells.
Why is multiparametric immunophenotyping superior to analyzing single markers?
Many cell surface markers are not entirely exclusive to one single cell type. Multiparametric immunophenotyping assesses multiple markers simultaneously, creating a complex “fingerprint” that allows for the highly precise identification of specific cell subsets, developmental stages, and abnormal disease states that a single marker could not definitively identify.
Glossary of Related Medical Terms
- Antigen: A molecular structure, often a protein or carbohydrate on a cell surface, that can be recognized by the immune system and specifically bound by an antibody.
- Cluster of Differentiation (CD): A standardized nomenclature system used to identify and investigate cell surface molecules providing targets for immunophenotyping of cells.
- Flow Cytometry: A laser-based technology used to analyze the physical and chemical characteristics of cells or particles suspended in a fluid as they pass through a beam of light.
- Fluorochrome: A fluorescent chemical compound conjugated to an antibody that re-emits light upon light excitation, used in flow cytometry to detect specific cell surface markers.
- Hematopoiesis: The biological process through which the body manufactures blood cells, originating from stem cells residing primarily in the bone marrow.
- Immunophenotyping: The process of using antibodies to identify cells based on the types of antigens or markers present on their surface.
- Lineage Negative (Lin-): A designation for cells, typically stem or progenitor cells, that lack the mature cell surface markers characteristic of fully differentiated blood cell lineages.
- Minimal Residual Disease (MRD): The small number of cancer cells that remain in the body during or after treatment, which do not cause physical signs or symptoms but can be detected via highly sensitive techniques like flow cytometry.
- Multipotent: The ability of a stem cell to develop into more than one cell type, but limited to the cell types within a closely related family or specific lineage.
Disclaimer: This article is intended for informational purposes only and is specifically targeted towards medical students. It is not intended to be a substitute for informed professional medical advice, diagnosis, or treatment. While the information presented here is derived from credible medical sources and is believed to be accurate and up-to-date, it is not guaranteed to be complete or error-free. See additional information.
References
- Majeti R, Park CY, Weissman IL. Identification of a hierarchy of multipotent hematopoietic progenitors in human cord blood. Cell Stem Cell. 2007;1(6):635-45.
- Qiao W, Wang W, Laurenti E, Turinsky AL, Wodak SJ, Bader GD, et al. Intercellular network structure and regulatory motifs in the human hematopoietic system. Mol Syst Biol. 2014;10:741.
- Wang X, Shok, J., Edinger, M., Warner, N., Bush-Donovan, C. Multiparametric Immunophenotyping of Human Hematopoietic Stem Cells and Progenitors Cells by Flow Cytometry. BD Biosciences Application Note. 2012:1-16.
- Weeda, V.; Mestrum, S.G.C.; Leers, M.P.G. Flow Cytometric Identification of Hematopoietic and Leukemic Blast Cells for Tailored Clinical Follow-Up of Acute Myeloid Leukemia. Int. J. Mol. Sci. 2022, 23, 10529. https://doi.org/10.3390/ijms231810529
- Hokland P, Hokland M, Daley J, Ritz J. Identification and cloning of a prethymic precursor T lymphocyte from a population of common acute lymphoblastic leukemia antigen (CALLA)-positive fetal bone marrow cells. J Exp Med. 1987;165(6):1749-54
- Hao QL, Zhu J, Price MA, Payne KJ, Barsky LW, Crooks GM. Identification of a novel, human multilymphoid progenitor in cord blood. Blood. 2001;97(12):3683-90.
- Edvardsson L, Dykes J, Olofsson T. Isolation and characterization of human myeloid progenitor populations–TpoR as discriminator between common myeloid and megakaryocyte/erythroid progenitors. Exp Hematol. 2006;34(5):599-609.
- Huang S, Chen Z, Yu JF, Young D, Bashey A, Ho AD, et al. Correlation between IL-3 receptor expression and growth potential of human CD34(+) hematopoietic cells from different tissues. Stem cells. 1999;17(5):265-72.
- O’Gorman MR, Gelman R. Inter- and intrainstitutional evaluation of automated volumetric capillary cytometry for the quantitation of CD4- and CD8-positive T lymphocytes in the peripheral blood of persons infected with human immunodeficiency virus. Site Investigators and the NIAID New CD4 Technologies Focus Group. Clin Diagn Lab Immunol. 1997;4(2):173-9.
- Romagnani S. Type 1 T helper and type 2 T helper cells: functions, regulation and role in protection and disease. Int J Clin Lab Res. 1991;21(2):152-8.
- Fontenot JD, Gavin MA, Rudensky AY. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat Immunol. 2003;4(4):330-6.
- Yu N, Li X, Song W, Li D, Yu D, Zeng X, et al. CD4(+)CD25 (+)CD127 (low/-) T cells: a more specific Treg population in human peripheral blood. Inflammation. 2012;35(6):1773-80.
- Mandelboim O, Malik P, Davis DM, Jo CH, Boyson JE, Strominger JL. Human CD16 as a lysis receptor mediating direct natural killer cell cytotoxicity. Proc Natl Acad Sci U S A. 1999;96(10):5640-4.
- Reeves RK, Gillis J, Wong FE, Yu Y, Connole M, Johnson RP. CD16- natural killer cells: enrichment in mucosal and secondary lymphoid tissues and altered function during chronic SIV infection. Blood. 2010;115(22):4439-46.
- Wang K, Wei G, Liu D. CD19: a biomarker for B cell development, lymphoma diagnosis and therapy. Exp Hematol Oncol. 2012;1(1):36.
- Carter RH, Wang Y, Brooks S. Role of CD19 signal transduction in B cell biology. Immunol Res. 2002;26(1-3):45-54.
- Caraux A, Klein B, Paiva B, Bret C, Schmitz A, Fuhler GM, et al. Circulating human B and plasma cells. Age-associated changes in counts and detailed characterization of circulating normal CD138- and CD138+ plasma cells. Haematologica. 2010;95(6):1016-20.
- Baez A, Alvarez-Laderas I, Piruat JI, Caballero-Velazquez T, Barbado MV, Millan-Ucles A, et al. The CD27 memory B cells display changes in the gene expression pattern in elderly individuals. Immunology. 2014;144(3):399-404.
- Pritz T, Lair J, Ban M, Keller M, Weinberger B, Krismer M, et al. Plasma cell numbers decrease in bone marrow of old patients. Eur J Immunol. 2015;45(3):738-46.
- Zamani F, Zare Shahneh F, Aghebati-Maleki L, Baradaran B. Induction of CD14 Expression and Differentiation to Monocytes or Mature Macrophages in Promyelocytic Cell Lines: New Approach. Adv Pharm Bull. 2013;3(2):329-32.
- Martin TR, Mongovin SM, Tobias PS, Mathison JC, Moriarty AM, Leturcq DJ, et al. The CD14 differentiation antigen mediates the development of endotoxin responsiveness during differentiation of mononuclear phagocytes. J Leukoc Biol. 1994;56(1):1-9.
- Passlick B, Flieger D, Ziegler-Heitbrock HW. Identification and characterization of a novel monocyte subpopulation in human peripheral blood. Blood. 1989;74(7):2527-34.
- Sulicka J, Surdacki A, Mikolajczyk T, Strach M, Gryglewska B, Cwiklinska M, et al. Elevated markers of inflammation and endothelial activation and increased counts of intermediate monocytes in adult survivors of childhood acute lymphoblastic leukemia. Immunobiology. 2013;218(5):810-6.
- Villadangos JA, Cardoso M, Steptoe RJ, van Berkel D, Pooley J, Carbone FR, et al. MHC class II expression is regulated in dendritic cells independently of invariant chain degradation. Immunity. 2001;14(6):739-49.
- Jongbloed SL, Kassianos AJ, McDonald KJ, Clark GJ, Ju X, Angel CE, et al. Human CD141+ (BDCA-3)+ dendritic cells (DCs) represent a unique myeloid DC subset that cross-presents necrotic cell antigens. J Exp Med. 2010;207(6):1247-60.
- Fujimoto H, Sakata T, Hamaguchi Y, Shiga S, Tohyama K, Ichiyama S, et al. Flow cytometric method for enumeration and classification of reactive immature granulocyte populations. Cytometry. 2000;42(6):371-8.
- Guerin E, Orabona M, Raquil MA, Giraudeau B, Bellier R, Gibot S, et al. Circulating immature granulocytes with T-cell killing functions predict sepsis deterioration*. Crit Care Med. 2014;42(9):2007-18.
- Mazzone A, Ricevuti G. Leukocyte CD11/CD18 integrins: biological and clinical relevance. Haematologica. 1995;80(2):161-75.
- Daley JM, Thomay AA, Connolly MD, Reichner JS, Albina JE. Use of Ly6G-specific monoclonal antibody to deplete neutrophils in mice. J Leukoc Biol. 2008;83(1):64-70.
- Yoon J, Terada A, Kita H. CD66b regulates adhesion and activation of human eosinophils. J Immunol. 2007;179(12):8454-62.
- Hennersdorf F, Florian S, Jakob A, Baumgartner K, Sonneck K, Nordheim A, et al. Identification of CD13, CD107a, and CD164 as novel basophil-activation markers and dissection of two response patterns in time kinetics of IgE-dependent upregulation. Cell Res. 2005;15(5):325-35.
- van Velzen JF, Laros-van Gorkom BAP, Pop GAM, van Heerde WL. Multicolor flow cytometry for evaluation of platelet surface antigens and activation markers. Thromb Res. 2012;130(1):92-8.
- Rogers CE, Bradley MS, Palsson BO, Koller MR. Flow cytometric analysis of human bone marrow perfusion cultures: erythroid development and relationship with burst-forming units-erythroid. Exp Hematol. 1996;24(5):597-604.
- Li W. Flow Cytometry in the Diagnosis of Leukemias. In: Li W, editor. Leukemia [Internet]. Brisbane (AU): Exon Publications; 2022 Oct 16. Chapter 4.
- Rix B, Maduro AH, Bridge KS and Grey W (2022) Markers for human haematopoietic stem cells: The disconnect between an identification marker and its function. Front. Physiol. 13:1009160. doi: 10.3389/fphys.2022.1009160
- Garg, S., Madkaikar, M., & Ghosh, K. (2013). Investigating cell surface markers on normal hematopoietic stem cells in three different niche conditions. International journal of stem cells, 6(2), 129–133. https://doi.org/10.15283/ijsc.2013.6.2.129
- Radu, P., Zurzu, M., Paic, V., Bratucu, M., Garofil, D., Tigora, A., Georgescu, V., Prunoiu, V., Pasnicu, C., Popa, F., Surlin, P., Surlin, V., & Strambu, V. (2023). CD34—Structure, Functions and Relationship with Cancer Stem Cells. Medicina, 59(5), 938. https://doi.org/10.3390/medicina59050938
- Sidney, L. E., Branch, M. J., Dunphy, S. E., Dua, H. S., & Hopkinson, A. (2014). Concise review: evidence for CD34 as a common marker for diverse progenitors. Stem cells (Dayton, Ohio), 32(6), 1380–1389. https://doi.org/10.1002/stem.1661