Procedure-At-A-Glance
- Assemble casting tray with a comb suitable to your needs.
- Combine the correct mass of agarose powder with TAE or TBE buffer in a microwave-safe flask (e.g., 1g in 100mL for a 1% gel).
- Microwave in short bursts, swirling gently, until the agarose is completely dissolved and the liquid is crystal clear. There must not be any small bubbles. Flask can get quite hot so please use oven mittens.
- Let the flask cool to touch (about 50°C). Add your DNA stain (e.g., EtBr or SYBR Safe) and swirl to mix.
- Pour the mixture into the assembled casting tray. Allow it to sit undisturbed at room temperature for 30–45 minutes until firm.
- Transfer the solidified gel (without the comb) into the electrophoresis tank and cover completely with running buffer.
- Mix your DNA samples with loading dye and carefully pipette them into the wells. Always include a DNA ladder in the first well.
- Connect the power supply (ensure the wells are at the negative/black electrode) and run at the appropriate voltage (e.g., 70-100 V) until the dye front has migrated sufficiently.
- Carefully remove the gel and view it under a UV transilluminator or gel documentation system.
Introduction
Unraveling the mysteries of life often requires peering into the microscopic world of DNA. Conventional Polymerase Chain Reaction (PCR) is a powerful technique that allows us to amplify specific DNA sequences, making them visible and easily analyzed for example in diagnosing specific beta-thalassemia mutations. We can use agarose gel to make sure that the right amplicons have been amplified before sending the PCR product for sequencing. This protocol focuses on the final step of this process: agarose gel electrophoresis, where we separate and visualize the amplified DNA fragments.
Imagine a tiny copy machine for DNA! PCR takes a target DNA sequence and replicates it millions of times through repeated cycles of heating, annealing and cooling. Specific primers flank the target region, guiding the DNA polymerase enzyme to faithfully copy the desired stretch. This exponential amplification allows us to detect even minute amounts of DNA, making PCR invaluable for various applications in medicine, research, and forensics.
But how do we see these amplified DNA fragments? This is where agarose gel electrophoresis comes in. Molten agarose, a seaweed-derived polymer, solidifies into a mesh-like network when cooled. When an electric field is applied, negatively charged DNA molecules migrate through the gel pores at different rates depending on their size. Smaller fragments move faster and travel farther, while larger ones are hindered by the gel matrix.
The gel is stained with a fluorescent nucleic acid dye like ethidium bromide (EtBr) or SYBR safe stains or RedSafeTM stain. These stains bind to the PCR amplicons and fluoresce when excited using ultraviolet (UV) light. Thus, allowing us to visualize these separated DNA bands. Each band represents a specific amplified DNA fragment, providing valuable information about the presence, size, and abundance of the target sequence in the original sample.
This protocol will guide you through the steps of preparing an agarose gel for conventional PCR analysis.
Principle
The foundation lies in the unique properties of agarose, a natural polymer extracted from seaweed. When heated, agarose forms a semi-solid, jelly-like matrix, riddled with tiny pores. These pores, like miniature tunnels, act as a sieving mechanism for molecules, especially charged ones like DNA.
DNA molecules, being negatively charged due to their phosphate backbone, experience an attractive force towards the positive electrode when an electric field is applied across the gel. This electrophoretic pull sets them on a journey through the agarose gel.
But not all DNA molecules navigate the gel with equal ease. This differential mobility, dictated by size, is the key to agarose gel's use. Smaller DNA fragments, due to their compact structure, can readily wiggle through the narrow pores, encountering less resistance and zipping towards the positive end. Larger fragments, on the other hand, face a tougher obstacle course. Their bulkier size makes them snag on the agarose strands, slowing down their progress.
Materials
For Agarose Gel
- Agarose powder
- 1x TBE or TAE buffer (Tris-Acetate-EDTA or Tris-Acetate-Borate buffer)
- Microwave
- Erlenmeyer flask or microwave-safe flask (e.g., 250 mL)
- Gel casting tray with comb
- Staining dye (e.g., ethidium bromide, SYBR Safe, GelRed)
- Gloves
- Safety goggles
For Sample Loading
- Micropipettes and pipette tips
- DNA samples (PCR products, plasmids, etc.)
- DNA loading dye
- DNA ladder (e.g., 100 bp DNA ladder)
Protocol
Preparation
- Wear gloves and safety goggles throughout the procedure.
- Weigh out the desired amount of agarose powder (e.g., 1 gram for a 1% gel in 100 mL buffer). Usually 1.5 - 2% gel is used. However, if the amplicons designed are very close together in size, 3% gel can be used.
- Add the agarose powder to the Erlenmeyer flask or microwave-safe flask.
- Pour in the TBE or TAE buffer until the desired volume is reached (e.g., 100 mL for a 1% gel). For less than 10 samples, a small gel of 30 mL can be made.
- Swirl gently to evenly distribute the agarose powder.
Melting
- Microwave the agarose/buffer mixture on high power for 30 - 60 seconds intervals, swirling the flask gently after each interval, until the agarose is completely dissolved and the solution is clear. The agarose gel is completely dissolved when there are no longer small, tiny bubbles in the solution. The solution should be clear without any bubbles when it settles.
- Be careful as the solution will be hot! Use heat-resistant gloves and handle the flask with caution.
Cooling and Pouring
- Assemble the tray, cassette and comb using the desired sizes of gel and number of wells.
- Add the desired amount of staining dye (e.g., 5 µL of ethidium bromide per 100 mL of agarose solution).
- Swirl gently to mix the dye.
- Carefully pour the warm agarose solution into the gel casting tray, ensuring the wells are oriented correctly. Ensure that there are no bubbles in the gel. Use a pipette tip to pop any bubbles or to push the bubbles to the side of the gel away from the path of wells.
- Allow the gel to solidify completely at room temperature for 30-45 minutes or until it's firm. The gels can be prepared in advance, wrapped in a cling film once solidified and placed in the fridge until ready to use. Do not prepare too far in advance as the gel may dry up if left too long in the fridge.
Sample Loading and Running
- Once the gel is solidified, carefully remove the comb.
- Place the gel cassette in the electrophoresis tank and fill it with enough buffer to cover the gel.
- Pipette your DNA samples (5 uL) including the non-template control, mixed with an equal amount of loading dye (if necessary), into the wells.
- Pipette the DNA ladder (3 uL), mixed with an equal amount of loading dye (if necessary) into one of the wells, preferably the first well.
- Close the lid of the tank and connect it to the power supply. Make sure that the top of the gel (where the samples are loaded) is on the negative side of the power supply as the DNA is negatively charged and will move towards the positive end.
- Run the electrophoresis at the desired voltage for the appropriate time (e.g., 70-100 V for 30-60 minutes).
Visualization
- After electrophoresis, transfer the gel to a UV transilluminator or gel documentation system and visualize the DNA bands or PCR amplicons.
- Take a picture or analyze the bands.
Interpretation
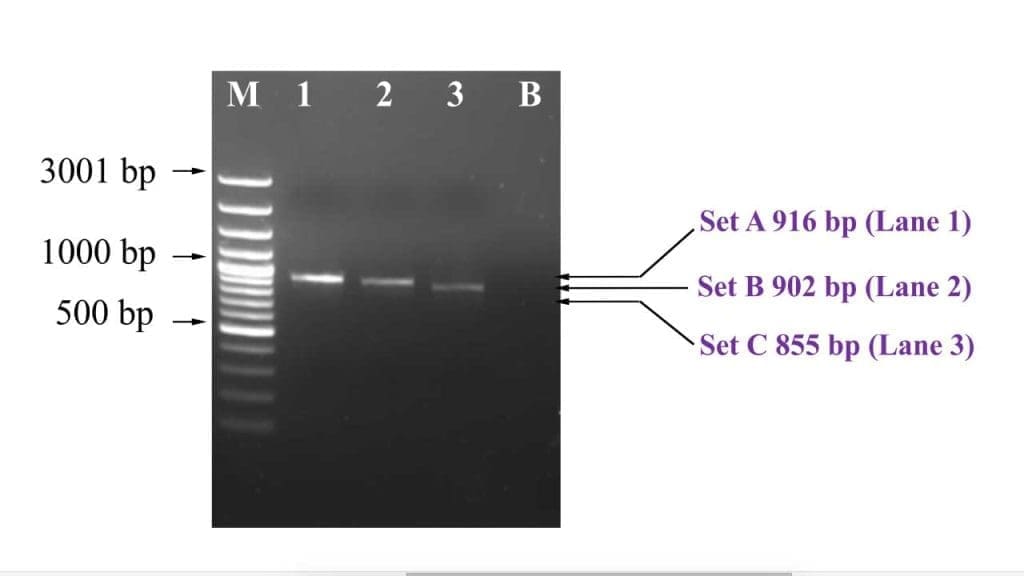
It is imperative to check that the right bands have been amplified especially using a DNA ladder. To be sure, the PCR amplicons can be sent for sequencing to ensure that the exact band that is required was correctly amplified. Non-template control is important as a negative control for contamination.
Using the right agarose concentration and the right well size goes a long way in reporting a PCR result.
Agarose Gel Preparation Tips
TAE vs TBE Buffers
Buffering Capacity and Stamina
TBE is the marathon runner of buffers. Because borate is a better buffer than acetate, TBE maintains its pH much longer during extended runs. If you are running a gel for several hours to separate very similar bands, TBE won't "exhaust" or allow the pH to drift. TAE, conversely, is prone to ion depletion; for very long runs, you might actually need to recirculate the buffer to keep the gradient steady.
Resolution and Fragment Size
If you are working with the small amplicons common in hematology PCR (like those for specific point mutations or small deletions), TBE is usually superior. It produces much sharper, tighter bands for fragments under 1,000 base pairs. However, for larger genomic DNA or large plasmids, TAE provides better separation and allows the DNA to move through the matrix with less "drag."
Downstream Compatibility
This is often the deciding factor in research. If you plan to cut a band out of the gel to use it for ligation, cloning, or specific sequencing protocols, TAE is the gold standard. The borate ions in TBE can stick to the DNA and inhibit certain enzymes (like ligases), which can ruin your next experiment. DNA extracted from a TAE gel is generally "cleaner" for enzymatic work.
TAE vs. TBE: Comparison at a Glance
| Feature | TAE (Tris-Acetate-EDTA) | TBE (Tris-Borate-EDTA) |
| Main Salt | Acetic Acid | Boric Acid |
| Buffering Capacity | Lower (can exhaust in long runs) | Higher (very stable over long periods) |
| DNA Migration | Faster migration | Slower migration |
| Resolution | Better for large fragments (>4 kb) | Better for small fragments (<1 kb) |
| Downstream Use | Ideal for DNA recovery/cloning | Borate may inhibit some enzymes |
| Heat Generation | Higher (at high voltage) | Lower (more conductive, stays cooler) |
Voltage Guidelines
Setting the voltage for your gel isn't just about picking a number on the power supply; it’s about balancing speed with the physical integrity of your samples. If the voltage is too low, your DNA will diffuse, leaving you with blurry, indistinct "blobs" instead of crisp bands. If it’s too high, the gel can overheat, causing your DNA to denature or, in extreme cases, turning your carefully prepared agarose back into a puddle of liquid.
The "Golden Rule": Volts per Centimeter
The most accurate way to determine voltage is not by the total voltage, but by the distance between the electrodes (the red and black terminals). The standard recommendation for most PCR applications is 5 to 10 V/cm.
To calculate the ideal voltage for your specific setup, use this formula:
Total Voltage = (Desired V/cm) x (Distance between electrodes in cm)
Example: If your electrophoresis tank has 15 cm between the two electrodes and you want to run it at a moderate 6 V/cm, you should set your power supply to 90V.
Low Voltage vs. High Voltage
| Setting | When to Use | The Result |
| Low (1–4 V/cm) | Large genomic DNA or complex mapping. | Higher resolution; very sharp bands but takes several hours (or overnight). |
| Medium (5–8 V/cm) | Standard PCR amplicons (e.g., beta-globin). | The "sweet spot" for most lab protocols; good balance of speed and clarity. |
| High (>10 V/cm) | Quick "yes/no" checks (e.g., is there DNA?). | Very fast (20 mins), but risks "smiling" bands and overheating the gel. |
Determining Gel Percentage
The percentage of agarose in your gel is essentially the "mesh size" of your molecular sieve. By adjusting the concentration of agarose, you control the size of the pores the DNA must wiggle through. Get the percentage wrong, and your DNA will either zip through too fast to separate or get stuck at the starting line like a car in a traffic jam.
The General Rule of Thumb
- High Percentage (2% – 3%): Creates a tight, dense matrix with small pores. This is ideal for separating small DNA fragments (50 bp – 1,000 bp).
- Low Percentage (0.7% – 1%): Creates a loose matrix with large pores. This is best for large DNA fragments (5 kb – 10 kb+).
Concentration Guidelines by Fragment Size
| Agarose Concentration (%) | Optimized Resolution Range (bp or kb) | Common Application |
| 0.7% | 5,000 – 10,000 bp (5–10 kb) | Genomic DNA, large plasmids |
| 1.0% | 500 – 5,000 bp | Standard PCR products, restriction digests |
| 1.5% | 200 – 3,000 bp | Small PCR amplicons, screening |
| 2.0% | 50 – 2,000 bp | High-resolution separation of small fragments |
| 3.0% | < 500 bp | Very small fragments, distinguishing between alleles |
How to Calculate the Mass Needed
To prepare your gel, you use a weight-to-volume percentage (w/v). The formula is:
Grams of Agarose = Desired %/100 x Volume of Buffer (mL)
Example: If you need a 2% gel and your casting tray holds 50 mL of buffer:
0.02 x 50 mL = 1 gram of agarose
Troubleshooting Tips for Agarose Gel Preparation
While agarose gel preparation seems straightforward, it can sometimes lead to frustration when things don't go as planned. Here are some common issues you might encounter and tips to get your gel back on track:
1. Gel not solidifying
- Possible cause: Wrong agarose powder concentration calculations or incomplete agarose melting or or insufficient cooling.
- Solution: Make sure that the agarose powder used is in the right concentration. Microwave the agarose solution in short bursts, swirling in between, until completely dissolved. Ensure the solution cools to around 50°C before pouring.
2. Bubbles in the gel
- Possible cause: Trapped air during pouring.
- Solution: Swirl the agarose gently while pouring and avoid over-heating the solution. You can also try poking the bubbles with a sterile pipette tip or push the bubbles to the side of the gel.
3. Bands blurry or smeared
- Possible cause: Overloading the wells, high voltage, or denatured DNA.
- Solution: Load smaller sample volumes, adjust the voltage based on your fragment size, and ensure proper sample preparation to avoid denaturation.
4. Faint or no bands:
- Possible cause: Low DNA concentration, insufficient staining, or inadequate electrophoresis time.
- Solution: Quantify your DNA samples, increase staining intensity, or run the gel for a longer time depending on your fragment size.
5. Uneven band migration:
- Possible cause: Tilted gel, uneven buffer levels, or bubbles under the comb.
- Solution: Ensure the gel tray is level, fill the tank with enough buffer to cover the gel evenly, and remove any bubbles trapped under the comb before electrophoresis.
6. The 'Smiling' Effect:
- Possible cause: f your bands look like a U-shape (the edges are higher than the center), your gel is running too hot. This usually happens at high voltages because the center of the gel dissipates heat more slowly than the edges.
- Solution: Lower the voltage.
4. Melting Gels:
- Possible cause: If you notice the buffer level dropping or the gel looking "warped" near the wells, your voltage is too high for the buffer capacity.
- Solution: TBE can handle slightly higher voltages than TAE without melting because it is more conductive.
Frequently Asked Questions (FAQs)
Frequently Asked Questions (FAQs)
How do I know what percentage of agarose gel to use?
The percentage of agarose determines the size of the pores in the gel matrix. Lower percentages (e.g., 0.7% - 1%) are used to resolve larger DNA fragments, while higher percentages (e.g., 2% - 3%) are better for separating smaller fragments, such as closely sized amplicons in hematological genetic testing.
Can I use plain water instead of TAE or TBE buffer to make and run the gel?
No. Plain water lacks the necessary ions to conduct an electrical current effectively, and it will not maintain the pH of the environment. Running a gel in water will cause the gel to melt from high resistance heat, and your DNA will not migrate properly.
Why are my DNA bands smearing instead of forming sharp lines?
Smeared bands can be caused by several factors: overloading the well with too much DNA, running the gel at a voltage that is too high (causing the gel to heat up), or using degraded/denatured DNA samples.
Is Ethidium Bromide safe to use on the open bench?
Ethidium Bromide is a known mutagen and potential carcinogen. It should always be handled in a designated area while wearing appropriate personal protective equipment (PPE), including nitrile gloves (not latex) and a lab coat. Many modern labs are switching to safer, less toxic alternatives like SYBR Safe or GelRed.
Why is my DNA running the wrong way (up instead of down)?
DNA is negatively charged due to its phosphate backbone. It will always migrate toward the positive (red) electrode. If your samples are running backward out of the wells, the electrodes on your power supply are plugged in backward. Always remember: "Run to Red."
Glossary of Related Medical & Scientific Terms
- Agarose: A highly purified polysaccharide extracted from seaweed that, when dissolved and cooled, forms a porous matrix ideal for separating nucleic acids.
- Amplicon: A piece of DNA or RNA that is the source and/or product of amplification or replication events, such as those produced by PCR.
- DNA Ladder: A solution containing DNA fragments of known lengths used as a reference to estimate the size of unknown DNA fragments in the sample wells.
- Ethidium Bromide (EtBr): A common fluorescent tag used in molecular biology laboratories for staining nucleic acids. It intercalates between DNA base pairs and fluoresces under UV light. (Note: It is a strong mutagen and must be handled with extreme care).
- Loading Dye: A colored, dense solution mixed with DNA samples before loading. It helps the sample sink into the well and provides a visible dye front to track the progress of the electrophoresis run.
- Polymerase Chain Reaction (PCR): A laboratory technique used to rapidly produce (amplify) millions to billions of copies of a specific segment of DNA.
Disclaimer: This protocol is intended for informational purposes only and may need to be modified depending on the specific laboratory procedures and patient circumstances. Always consult with a qualified healthcare professional for guidance. See additional information.
References
- https://www.addgene.org/protocols/gel-electrophoresis/
- R. Green MR, Sambrook J. Molecular Cloning: A Laboratory Manual (Fourth Edition), Volume 1, 2 & 3. Cold Spring Harbor Laboratory Press. 2012.
- Armstrong JA, Schulz JR. Agarose Gel Electrophoresis. 2015. Current Protocols: Essential Laboratory Techniques. https://doi.org/10.1002/9780470089941.et0702s10